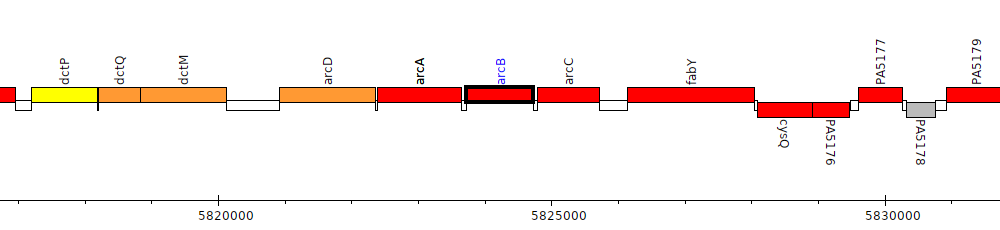
Pseudomonas aeruginosa PAO1, PA5172 (arcB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019546 | arginine deiminase pathway | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
6438064 | Reviewed by curator |
| Molecular Function | GO:0004585 | ornithine carbamoyltransferase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
6438064 | Reviewed by curator |
| Molecular Function | GO:0004585 | ornithine carbamoyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01109
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.1370
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016597 | amino acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.1370
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016743 | carboxyl- or carbamoyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00658
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006591 | ornithine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01109
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Arginine and proline metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Urea cycle and metabolism of amino groups |
ECO:0000037
not_recorded |
|||
| KEGG | pae00220 | Arginine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00102 | Ornithine carbamoyltransferase signature | IPR002292 | Ornithine/putrescine carbamoyltransferase | 312 | 323 | 1.0E-30 |
| PRINTS | PR00100 | Aspartate/ornithine carbamoyltransferase superfamily signature | IPR006130 | Aspartate/ornithine carbamoyltransferase | 298 | 321 | 3.2E-25 |
| Gene3D | G3DSA:3.40.50.1370 | Aspartate/ornithine carbamoyltransferase | IPR036901 | Aspartate/ornithine carbamoyltransferase superfamily | 152 | 320 | 0.0 |
| PRINTS | PR00102 | Ornithine carbamoyltransferase signature | IPR002292 | Ornithine/putrescine carbamoyltransferase | 124 | 138 | 1.0E-30 |
| PRINTS | PR00100 | Aspartate/ornithine carbamoyltransferase superfamily signature | IPR006130 | Aspartate/ornithine carbamoyltransferase | 267 | 276 | 3.2E-25 |
| Gene3D | G3DSA:3.40.50.1370 | Aspartate/ornithine carbamoyltransferase | IPR036901 | Aspartate/ornithine carbamoyltransferase superfamily | 5 | 333 | 0.0 |
| Hamap | MF_01109 | Ornithine carbamoyltransferase, catabolic [argI]. | IPR024904 | Ornithine carbamoyltransferase | 7 | 334 | 81.382256 |
| Pfam | PF00185 | Aspartate/ornithine carbamoyltransferase, Asp/Orn binding domain | IPR006131 | Aspartate/ornithine carbamoyltransferase, Asp/Orn-binding domain | 156 | 331 | 6.7E-55 |
| PANTHER | PTHR45753 | ORNITHINE CARBAMOYLTRANSFERASE, MITOCHONDRIAL | - | - | 3 | 333 | 2.8E-101 |
| FunFam | G3DSA:3.40.50.1370:FF:000003 | Ornithine carbamoyltransferase | - | - | 5 | 162 | 8.9E-83 |
| PRINTS | PR00100 | Aspartate/ornithine carbamoyltransferase superfamily signature | IPR006130 | Aspartate/ornithine carbamoyltransferase | 135 | 146 | 3.2E-25 |
| SUPERFAMILY | SSF53671 | Aspartate/ornithine carbamoyltransferase | IPR036901 | Aspartate/ornithine carbamoyltransferase superfamily | 4 | 334 | 5.76E-100 |
| PRINTS | PR00102 | Ornithine carbamoyltransferase signature | IPR002292 | Ornithine/putrescine carbamoyltransferase | 51 | 65 | 1.0E-30 |
| NCBIfam | TIGR00658 | JCVI: ornithine carbamoyltransferase | IPR002292 | Ornithine/putrescine carbamoyltransferase | 8 | 333 | 0.0 |
| PRINTS | PR00100 | Aspartate/ornithine carbamoyltransferase superfamily signature | IPR006130 | Aspartate/ornithine carbamoyltransferase | 53 | 72 | 3.2E-25 |
| PRINTS | PR00102 | Ornithine carbamoyltransferase signature | IPR002292 | Ornithine/putrescine carbamoyltransferase | 84 | 97 | 1.0E-30 |
| Pfam | PF02729 | Aspartate/ornithine carbamoyltransferase, carbamoyl-P binding domain | IPR006132 | Aspartate/ornithine carbamoyltransferase, carbamoyl-P binding | 8 | 148 | 1.0E-45 |
| PRINTS | PR00102 | Ornithine carbamoyltransferase signature | IPR002292 | Ornithine/putrescine carbamoyltransferase | 229 | 239 | 1.0E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.