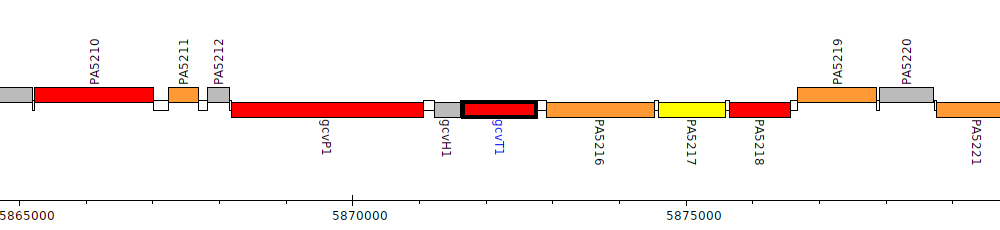
Pseudomonas aeruginosa PAO1, PA5215 (gcvT1)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019464 | glycine decarboxylation via glycine cleavage system | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
3023185 | Reviewed by curator |
| Molecular Function | GO:0004047 | aminomethyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00259
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006546 | glycine catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00259
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1360.120
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | One carbon pool by folate |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00670 | One carbon pool by folate | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glycine, serine and threonine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Nitrogen metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.30.70.1400:FF:000001 | Aminomethyltransferase | - | - | 51 | 137 | 9.3E-27 |
| SUPERFAMILY | SSF101790 | Aminomethyltransferase beta-barrel domain | IPR029043 | Glycine cleavage T-protein/YgfZ, C-terminal | 269 | 357 | 9.81E-26 |
| PIRSF | PIRSF006487 | GCST | - | - | 1 | 358 | 4.4E-80 |
| FunFam | G3DSA:2.40.30.110:FF:000001 | Aminomethyltransferase | - | - | 277 | 360 | 4.0E-38 |
| NCBIfam | TIGR00528 | JCVI: glycine cleavage system aminomethyltransferase GcvT | IPR006223 | Glycine cleavage system T protein | 4 | 354 | 1.0E-124 |
| Gene3D | G3DSA:3.30.70.1400 | - | - | - | 51 | 137 | 1.9E-84 |
| Hamap | MF_00259 | Aminomethyltransferase [gcvT]. | IPR022903 | Glycine cleavage system T protein, bacteria | 1 | 355 | 43.610336 |
| PANTHER | PTHR43757 | AMINOMETHYLTRANSFERASE | IPR028896 | Aminomethyltransferase-like | 4 | 356 | 2.7E-96 |
| Pfam | PF01571 | Aminomethyltransferase folate-binding domain | IPR006222 | Aminomethyltransferase, folate-binding domain | 7 | 253 | 7.3E-80 |
| Pfam | PF08669 | Glycine cleavage T-protein C-terminal barrel domain | IPR013977 | Glycine cleavage T-protein, C-terminal barrel domain | 280 | 353 | 1.7E-15 |
| Gene3D | G3DSA:2.40.30.110 | - | - | - | 277 | 360 | 1.2E-33 |
| Gene3D | G3DSA:3.30.1360.120 | Probable tRNA modification gtpase trme; domain 1 | IPR027266 | GTP-binding protein TrmE/Aminomethyltransferase GcvT, domain 1 | 5 | 230 | 1.9E-84 |
| Gene3D | G3DSA:4.10.1250.10 | Aminomethyltransferase fragment | - | - | 231 | 274 | 7.3E-26 |
| FunFam | G3DSA:4.10.1250.10:FF:000001 | Aminomethyltransferase | - | - | 231 | 275 | 5.2E-21 |
| SUPERFAMILY | SSF103025 | Folate-binding domain | - | - | 4 | 276 | 4.55E-87 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.