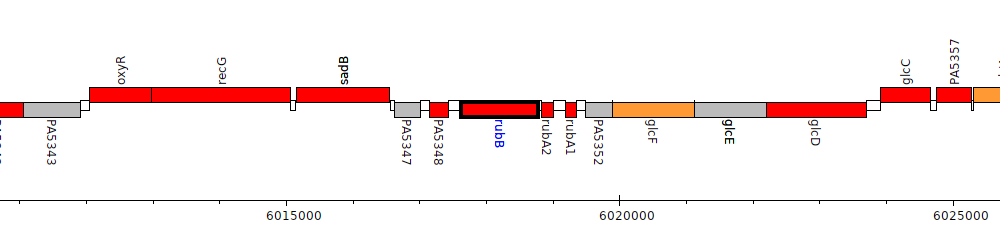
Pseudomonas aeruginosa PAO1, PA5349 (rubB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0043448 | alkane catabolic process | Inferred from Genetic Interaction | ECO:0000316 genetic interaction evidence used in manual assertion |
14574114 | Reviewed by curator |
| Molecular Function | GO:0015045 | rubredoxin-NAD(P)+ reductase activity | Inferred from Genetic Interaction | ECO:0000316 genetic interaction evidence used in manual assertion |
14574114 | Reviewed by curator |
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | P221-PWY | octane oxidation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 147 | 165 | 6.6E-22 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 233 | 247 | 4.1E-15 |
| FunFam | G3DSA:3.50.50.60:FF:000319 | Rubredoxin-NAD(+) reductase | - | - | 110 | 244 | 2.0E-70 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 359 | 374 | 4.1E-15 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 110 | 244 | 1.0E-95 |
| Pfam | PF18113 | Rubredoxin binding C-terminal domain | IPR041364 | Rubredoxin binding domain | 309 | 379 | 2.8E-24 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 257 | 279 | 6.6E-22 |
| Gene3D | G3DSA:3.30.390.120 | - | - | - | 316 | 379 | 1.1E-25 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 7 | 281 | 5.6E-51 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 8 | 280 | 1.0E-95 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 323 | 5.4E-36 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 102 | 120 | 6.6E-22 |
| FunFam | G3DSA:3.30.390.120:FF:000002 | Rubredoxin-NAD(+) reductase | - | - | 316 | 379 | 8.9E-33 |
| PANTHER | PTHR43429 | PYRIDINE NUCLEOTIDE-DISULFIDE OXIDOREDUCTASE DOMAIN-CONTAINING | - | - | 6 | 308 | 7.2E-60 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 232 | 248 | 6.6E-22 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 272 | 279 | 4.1E-15 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 6 | 28 | 4.1E-15 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 7 | 26 | 6.6E-22 |
| PRINTS | PR00411 | Pyridine nucleotide disulphide reductase class-I signature | - | - | 147 | 172 | 4.1E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.