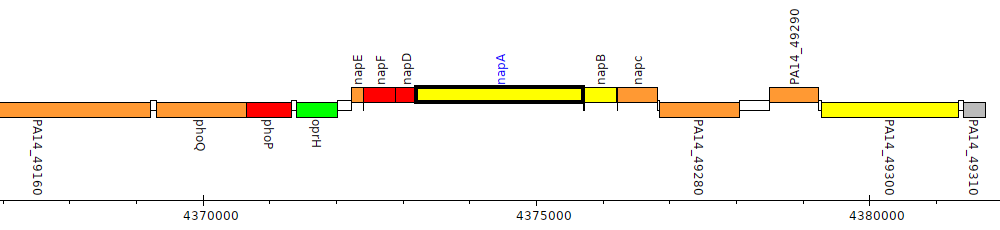
Pseudomonas aeruginosa UCBPP-PA14, PA14_49250 (napA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0030151 | molybdenum ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01630
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0043546 | molybdopterin cofactor binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01568
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04879
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008940 | nitrate reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01630
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Nitrogen metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pau01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR01706 | JCVI: periplasmic nitrate reductase subunit alpha | IPR010051 | Periplasmic nitrate reductase, large subunit | 3 | 829 | 0.0 |
| PANTHER | PTHR43105 | RESPIRATORY NITRATE REDUCTASE | - | - | 42 | 830 | 0.0 |
| Hamap | MF_01630 | Periplasmic nitrate reductase [napA]. | IPR010051 | Periplasmic nitrate reductase, large subunit | 1 | 831 | 32.362389 |
| FunFam | G3DSA:2.40.40.20:FF:000005 | Periplasmic nitrate reductase | - | - | 707 | 831 | 2.0E-52 |
| Gene3D | G3DSA:3.40.50.740 | - | - | - | 98 | 626 | 0.0 |
| Pfam | PF01568 | Molydopterin dinucleotide binding domain | IPR006657 | Molybdopterin dinucleotide-binding domain | 717 | 825 | 1.2E-24 |
| Gene3D | G3DSA:2.40.40.20 | - | - | - | 707 | 830 | 3.8E-53 |
| SUPERFAMILY | SSF53706 | Formate dehydrogenase/DMSO reductase, domains 1-3 | - | - | 37 | 648 | 0.0 |
| Pfam | PF04879 | Molybdopterin oxidoreductase Fe4S4 domain | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 43 | 94 | 1.4E-14 |
| CDD | cd02754 | MopB_Nitrate-R-NapA-like | - | - | 45 | 707 | 0.0 |
| Gene3D | G3DSA:3.40.228.10 | Dimethylsulfoxide Reductase, domain 2 | - | - | 177 | 706 | 0.0 |
| SMART | SM00926 | Molybdop_Fe4S4_2 | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 41 | 95 | 4.3E-9 |
| CDD | cd02791 | MopB_CT_Nitrate-R-NapA-like | IPR041957 | Nitrate reductase NapA-like, molybdopterin-binding domain | 714 | 829 | 9.28884E-54 |
| Gene3D | G3DSA:3.30.200.210 | - | - | - | 42 | 669 | 0.0 |
| Pfam | PF00384 | Molybdopterin oxidoreductase | IPR006656 | Molybdopterin oxidoreductase | 98 | 571 | 3.9E-86 |
| SUPERFAMILY | SSF50692 | ADC-like | IPR009010 | Aspartate decarboxylase-like domain superfamily | 690 | 829 | 4.14E-42 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.