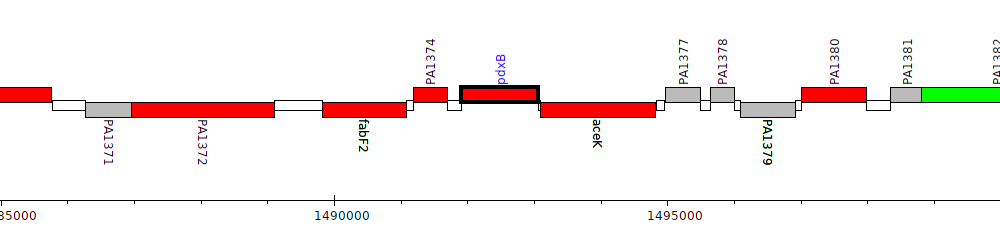
Pseudomonas aeruginosa PAO1, PA1375 (pdxB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008615 | pyridoxine biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2681152 | Reviewed by curator |
| Molecular Function | GO:0051287 | NAD binding | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2681152 | Reviewed by curator |
| Biological Process | GO:0042823 | pyridoxal phosphate biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2681152 | Reviewed by curator |
| Molecular Function | GO:0033711 | 4-phosphoerythronate dehydrogenase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
2681152 | Reviewed by curator |
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00389
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008615 | pyridoxine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01825
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02826
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046983 | protein dimerization activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF11890
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0033711 | 4-phosphoerythronate dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01825
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01825
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Butanoate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Vitamin B6 metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00750 | Vitamin B6 metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glycine, serine and threonine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PYRIDOXSYN-PWY | pyridoxal 5'-phosphate biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Ascorbate and aldarate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Phenylalanine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Fructose and mannose metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Galactose metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Lysine biosynthesis |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Nucleotide sugars metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF11890 | Domain of unknown function (DUF3410) | IPR024531 | Erythronate-4-phosphate dehydrogenase, dimerisation domain | 289 | 376 | 2.5E-22 |
| CDD | cd12158 | ErythrP_dh | IPR020921 | Erythronate-4-phosphate dehydrogenase | 2 | 350 | 0.0 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 1 | 277 | 2.2E-118 |
| Pfam | PF02826 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD binding domain | IPR006140 | D-isomer specific 2-hydroxyacid dehydrogenase, NAD-binding domain | 107 | 256 | 7.6E-35 |
| Hamap | MF_01825 | Erythronate-4-phosphate dehydrogenase [pdxB]. | IPR020921 | Erythronate-4-phosphate dehydrogenase | 1 | 363 | 44.927265 |
| FunFam | G3DSA:3.40.50.720:FF:000093 | Erythronate-4-phosphate dehydrogenase | - | - | 92 | 257 | 2.5E-75 |
| Pfam | PF00389 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | IPR006139 | D-isomer specific 2-hydroxyacid dehydrogenase, catalytic domain | 33 | 277 | 4.5E-15 |
| Gene3D | G3DSA:3.30.1370.170 | - | IPR038251 | PdxB, dimerisation domain superfamily | 293 | 380 | 6.7E-29 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 92 | 257 | 2.2E-118 |
| SUPERFAMILY | SSF52283 | Formate/glycerate dehydrogenase catalytic domain-like | - | - | 5 | 103 | 1.36E-15 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 91 | 258 | 5.56E-37 |
| PANTHER | PTHR42938 | FORMATE DEHYDROGENASE 1 | - | - | 33 | 274 | 1.2E-36 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.