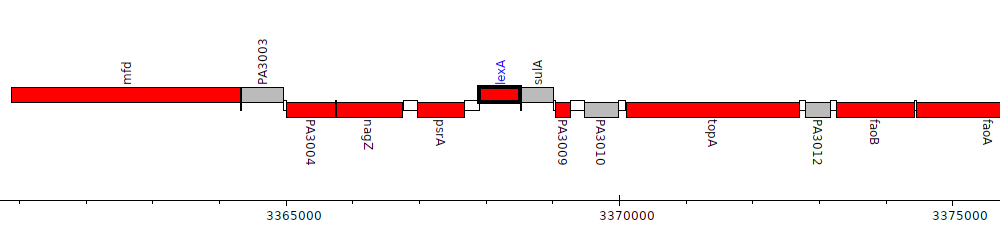
Pseudomonas aeruginosa PAO1, PA3007 (lexA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0009432 | SOS response | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
7027255 | Reviewed by curator |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
1494343 | Reviewed by curator |
| Biological Process | GO:0050896 | response to stimulus | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0009432 | SOS response |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00015
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PR00726
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0045892 | negative regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00015
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR00726
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004252 | serine-type endopeptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00015
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:PF01726
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.10.109.10 | Umud Fragment, subunit A | - | - | 81 | 204 | 2.0E-38 |
| SUPERFAMILY | SSF51306 | LexA/Signal peptidase | IPR036286 | LexA/Signal peptidase-like superfamily | 82 | 203 | 6.8E-36 |
| PANTHER | PTHR33516 | LEXA REPRESSOR | - | - | 1 | 203 | 5.8E-56 |
| CDD | cd06529 | S24_LexA-like | IPR039418 | LexA-like | 117 | 193 | 2.14595E-21 |
| Hamap | MF_00015 | LexA repressor [lexA]. | IPR006200 | Transcription regulator LexA | 3 | 204 | 32.413368 |
| FunFam | G3DSA:1.10.10.10:FF:000009 | LexA repressor | - | - | 1 | 69 | 2.6E-33 |
| PRINTS | PR00726 | Repressor LexA serine protease (S24) family signature | IPR006197 | Peptidase S24, LexA-like | 128 | 139 | 1.9E-12 |
| PRINTS | PR00726 | Repressor LexA serine protease (S24) family signature | IPR006197 | Peptidase S24, LexA-like | 117 | 127 | 1.9E-12 |
| Pfam | PF01726 | LexA DNA binding domain | IPR006199 | LexA repressor, DNA-binding domain | 1 | 65 | 2.0E-31 |
| NCBIfam | TIGR00498 | JCVI: transcriptional repressor LexA | IPR006200 | Transcription regulator LexA | 1 | 204 | 2.2E-74 |
| FunFam | G3DSA:2.10.109.10:FF:000001 | LexA repressor | - | - | 81 | 204 | 2.4E-53 |
| PRINTS | PR00726 | Repressor LexA serine protease (S24) family signature | IPR006197 | Peptidase S24, LexA-like | 156 | 168 | 1.9E-12 |
| SUPERFAMILY | SSF46785 | Winged helix DNA-binding domain | IPR036390 | Winged helix DNA-binding domain superfamily | 3 | 70 | 1.7E-21 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 1 | 68 | 1.2E-25 |
| Pfam | PF00717 | Peptidase S24-like | IPR015927 | Peptidase S24/S26A/S26B/S26C | 83 | 196 | 1.2E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.