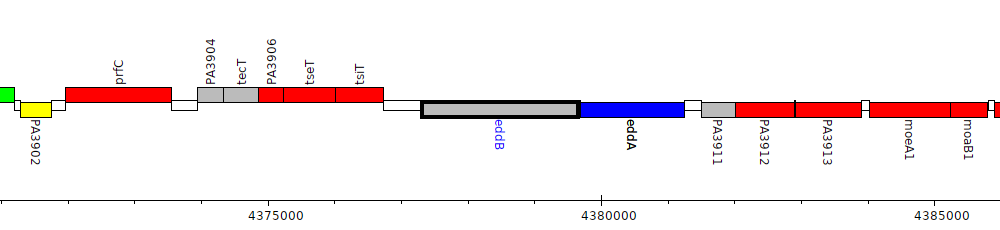
Pseudomonas aeruginosa PAO1, PA3909 (eddB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006308 | DNA catabolic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
20370819 | Reviewed by curator |
| Molecular Function | GO:0004536 | deoxyribonuclease activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20370819 | Reviewed by curator |
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF03372
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00932 | Lamin Tail Domain | IPR001322 | Lamin tail domain | 21 | 129 | 3.1E-15 |
| Gene3D | G3DSA:3.60.10.10 | Endonuclease/exonuclease/phosphatase | IPR036691 | Endonuclease/exonuclease/phosphatase superfamily | 454 | 767 | 2.3E-36 |
| CDD | cd10283 | MnuA_DNase1-like | - | - | 454 | 767 | 3.62249E-52 |
| Pfam | PF03372 | Endonuclease/Exonuclease/phosphatase family | IPR005135 | Endonuclease/exonuclease/phosphatase | 457 | 760 | 5.6E-8 |
| NCBIfam | NF033681 | NCBIFAM: ExeM/NucH family extracellular endonuclease | IPR047971 | Extracellular endonuclease ExeM-like | 221 | 769 | 0.0 |
| PANTHER | PTHR42834 | ENDONUCLEASE/EXONUCLEASE/PHOSPHATASE FAMILY PROTEIN (AFU_ORTHOLOGUE AFUA_3G09210) | - | - | 195 | 769 | 0.0 |
| FunFam | G3DSA:3.60.10.10:FF:000072 | Extracellular nuclease | - | - | 454 | 767 | 0.0 |
| SUPERFAMILY | SSF56219 | DNase I-like | IPR036691 | Endonuclease/exonuclease/phosphatase superfamily | 452 | 768 | 6.62E-36 |
| CDD | cd04486 | YhcR_OBF_like | - | - | 221 | 301 | 2.73461E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.