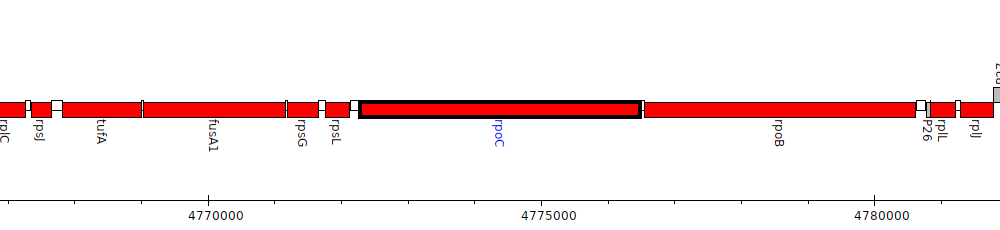
Pseudomonas aeruginosa PAO1, PA4269 (rpoC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006351 | transcription, DNA-templated | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
6287430 | Reviewed by curator |
| Biological Process | GO:0006351 | transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00623
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003899 | DNA-directed 5'-3' RNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00623
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00623
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Purine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Pyrimidine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae03020 | RNA polymerase | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:1.10.132.30:FF:000003 | DNA-directed RNA polymerase subunit beta | - | - | 638 | 790 | 1.1E-76 |
| FunFam | G3DSA:4.10.860.120:FF:000001 | DNA-directed RNA polymerase subunit beta | - | - | 12 | 132 | 5.5E-69 |
| Hamap | MF_01322 | DNA-directed RNA polymerase subunit beta' [rpoC]. | IPR012754 | DNA-directed RNA polymerase, subunit beta-prime, bacterial type | 14 | 1387 | 19.055861 |
| Coils | Coil | Coil | - | - | 191 | 211 | - |
| Gene3D | G3DSA:4.10.860.120 | RNA polymerase II, clamp domain | IPR044893 | RNA polymerase Rpb1, clamp domain superfamily | 8 | 142 | 1.5E-24 |
| Gene3D | G3DSA:2.40.50.100 | - | - | - | 948 | 1022 | 1.1E-24 |
| Gene3D | G3DSA:2.40.40.20 | - | - | - | 332 | 486 | 5.6E-48 |
| Gene3D | G3DSA:2.40.50.100 | - | - | - | 1151 | 1215 | 2.7E-68 |
| SUPERFAMILY | SSF64484 | beta and beta-prime subunits of DNA dependent RNA-polymerase | - | - | 16 | 1363 | 0.0 |
| Pfam | PF00623 | RNA polymerase Rpb1, domain 2 | IPR000722 | RNA polymerase, alpha subunit | 389 | 485 | 2.7E-28 |
| CDD | cd01609 | RNAP_beta'_N | - | - | 15 | 813 | 0.0 |
| Gene3D | G3DSA:2.40.50.100 | - | - | - | 1023 | 1126 | 3.9E-33 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1367 | 1399 | - |
| Pfam | PF04998 | RNA polymerase Rpb1, domain 5 | IPR007081 | RNA polymerase Rpb1, domain 5 | 766 | 1316 | 2.8E-77 |
| SMART | SM00663 | rpolaneu7 | IPR006592 | RNA polymerase, N-terminal | 235 | 514 | 0.0 |
| Gene3D | G3DSA:1.10.40.90 | - | - | - | 370 | 413 | 2.4E-21 |
| PANTHER | PTHR19376 | DNA-DIRECTED RNA POLYMERASE | IPR045867 | DNA-directed RNA polymerase, subunit beta-prime | 23 | 1363 | 0.0 |
| Gene3D | G3DSA:1.10.274.100 | RNA polymerase Rpb1, domain 3 | IPR042102 | RNA polymerase Rpb1, domain 3 superfamily | 489 | 637 | 4.3E-44 |
| Pfam | PF04983 | RNA polymerase Rpb1, domain 3 | IPR007066 | RNA polymerase Rpb1, domain 3 | 489 | 643 | 5.7E-30 |
| Gene3D | G3DSA:1.10.1790.20 | - | - | - | 1135 | 1314 | 2.7E-68 |
| CDD | cd02655 | RNAP_beta'_C | - | - | 907 | 1363 | 8.21481E-83 |
| NCBIfam | TIGR02386 | JCVI: DNA-directed RNA polymerase subunit beta' | IPR012754 | DNA-directed RNA polymerase, subunit beta-prime, bacterial type | 19 | 1366 | 0.0 |
| Pfam | PF00623 | RNA polymerase Rpb1, domain 2 | IPR000722 | RNA polymerase, alpha subunit | 344 | 381 | 2.4E-7 |
| Pfam | PF05000 | RNA polymerase Rpb1, domain 4 | IPR007083 | RNA polymerase Rpb1, domain 4 | 674 | 763 | 3.4E-20 |
| Gene3D | G3DSA:1.10.132.30 | - | IPR038120 | RNA polymerase Rpb1, funnel domain superfamily | 638 | 790 | 7.7E-68 |
| Gene3D | G3DSA:1.10.150.390 | - | - | - | 1316 | 1376 | 6.1E-27 |
| FunFam | G3DSA:1.10.40.90:FF:000001 | DNA-directed RNA polymerase subunit beta | - | - | 372 | 413 | 3.5E-22 |
| FunFam | G3DSA:1.10.150.390:FF:000002 | DNA-directed RNA polymerase subunit beta'' | - | - | 1316 | 1376 | 1.3E-31 |
| Pfam | PF04997 | RNA polymerase Rpb1, domain 1 | IPR007080 | RNA polymerase Rpb1, domain 1 | 16 | 342 | 2.6E-90 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.