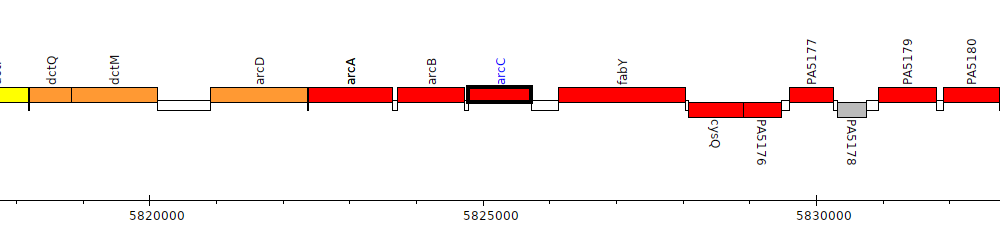
Pseudomonas aeruginosa PAO1, PA5173 (arcC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019546 | arginine deiminase pathway | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
6438064 | Reviewed by curator |
| Molecular Function | GO:0008804 | carbamate kinase activity | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
6438064 | Reviewed by curator |
| Molecular Function | GO:0008804 | carbamate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd04235
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006525 | arginine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd04235
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | CITRULLINE-DEG-PWY | L-citrulline degradation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00220 | Arginine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glutamate metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | ARGDEGRAD-PWY | L-arginine degradation V (arginine deiminase pathway) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Arginine and proline metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY0-41 | allantoin degradation IV (anaerobic) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Nitrogen metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 103 | 122 | 2.6E-70 |
| PIRSF | PIRSF000723 | Carbamate_kin | IPR003964 | Carbamate kinase | 1 | 302 | 9.4E-112 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 71 | 89 | 2.6E-70 |
| CDD | cd04235 | AAK_CK | IPR003964 | Carbamate kinase | 2 | 301 | 0.0 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 257 | 272 | 2.6E-70 |
| Gene3D | G3DSA:3.40.1160.10 | - | IPR036393 | Acetylglutamate kinase-like superfamily | 1 | 301 | 1.2E-106 |
| FunFam | G3DSA:3.40.1160.10:FF:000007 | Carbamate kinase | - | - | 1 | 302 | 2.1E-128 |
| NCBIfam | TIGR00746 | JCVI: carbamate kinase | IPR003964 | Carbamate kinase | 1 | 301 | 0.0 |
| SUPERFAMILY | SSF53633 | Carbamate kinase-like | IPR036393 | Acetylglutamate kinase-like superfamily | 1 | 300 | 3.8E-78 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 174 | 192 | 2.6E-70 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 151 | 170 | 2.6E-70 |
| PANTHER | PTHR30409 | CARBAMATE KINASE | IPR003964 | Carbamate kinase | 2 | 302 | 5.8E-124 |
| Pfam | PF00696 | Amino acid kinase family | IPR001048 | Aspartate/glutamate/uridylate kinase | 1 | 283 | 1.3E-25 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 206 | 221 | 2.6E-70 |
| PRINTS | PR01469 | Bacterial carbamate kinase signature | - | - | 41 | 60 | 2.6E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.