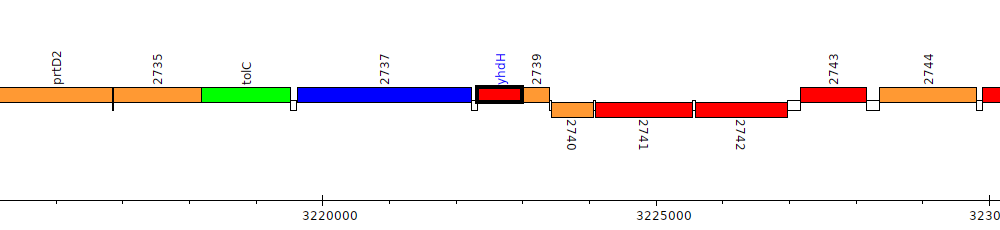
Pseudomonas fluorescens F113, PSF113_2738 (yhdH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pfe01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfe00640 | Propanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43677 | SHORT-CHAIN DEHYDROGENASE/REDUCTASE | - | - | 3 | 220 | 4.9E-24 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 23 | 187 | 2.8E-67 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 14 | 185 | 9.51E-36 |
| Pfam | PF00107 | Zinc-binding dehydrogenase | IPR013149 | Alcohol dehydrogenase-like, C-terminal | 56 | 170 | 3.2E-13 |
| Gene3D | G3DSA:3.90.180.10 | - | - | - | 2 | 219 | 2.8E-67 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.