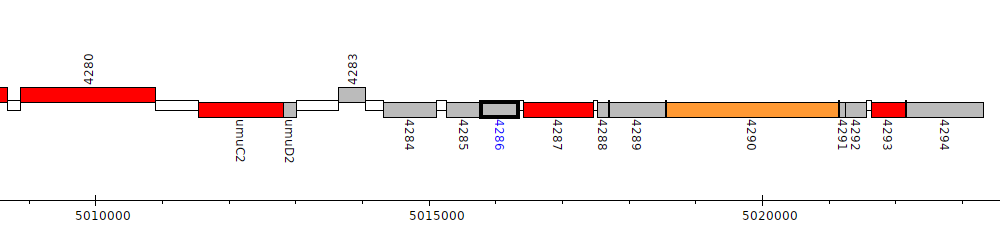
Pseudomonas fluorescens F113, PSF113_4286
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0016998 | cell wall macromolecule catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00182
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004568 | chitinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00182
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006032 | chitin catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00182
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | chitin degradation II (Vibrio) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | chitin degradation III (Serratia) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | chitin degradation I (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR34408 | FAMILY PROTEIN, PUTATIVE-RELATED | - | - | 3 | 182 | 1.1E-17 |
| CDD | cd00325 | chitinase_GH19 | - | - | 3 | 165 | 4.07981E-22 |
| SUPERFAMILY | SSF53955 | Lysozyme-like | IPR023346 | Lysozyme-like domain superfamily | 2 | 185 | 4.5E-47 |
| Pfam | PF00182 | Chitinase class I | IPR000726 | Glycoside hydrolase, family 19, catalytic | 39 | 142 | 6.0E-6 |
| Gene3D | G3DSA:1.10.530.10 | - | - | - | 3 | 183 | 2.7E-38 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.