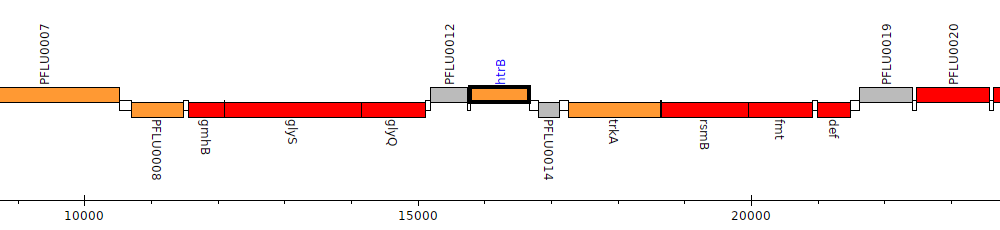
Pseudomonas fluorescens SBW25, PFLU0013 (htrB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016740 | transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd07984
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:cd07984
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00604 | Glycosphingolipid biosynthesis - ganglio series | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00253 | Tetracycline biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00950 | Isoquinoline alkaloid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00626 | Naphthalene degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00680 | Methane metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00540 | Lipopolysaccharide biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00360 | Phenylalanine metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00940 | Phenylpropanoid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00564 | Glycerophospholipid metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00310 | Lysine degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00642 | Ethylbenzene degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00960 | Tropane, piperidine and pyridine alkaloid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00904 | Diterpenoid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00623 | Toluene degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00650 | Butanoate metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00906 | Carotenoid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00903 | Limonene and pinene degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00061 | Fatty acid biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00350 | Tyrosine metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00330 | Arginine and proline metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00965 | Betalain biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00942 | Anthocyanin biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00340 | Histidine metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00565 | Ether lipid metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00362 | Benzoate degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00627 | Aminobenzoate degradation | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF03279 | Bacterial lipid A biosynthesis acyltransferase | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 8 | 286 | 1.8E-46 |
| PIRSF | PIRSF026649 | MsbB | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 1 | 295 | 4.4E-95 |
| CDD | cd07984 | LPLAT_LABLAT-like | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 97 | 285 | 4.01439E-43 |
| PANTHER | PTHR30606 | LIPID A BIOSYNTHESIS LAUROYL ACYLTRANSFERASE | IPR004960 | Bacterial lipid A biosynthesis acyltransferase | 5 | 293 | 1.3E-51 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.