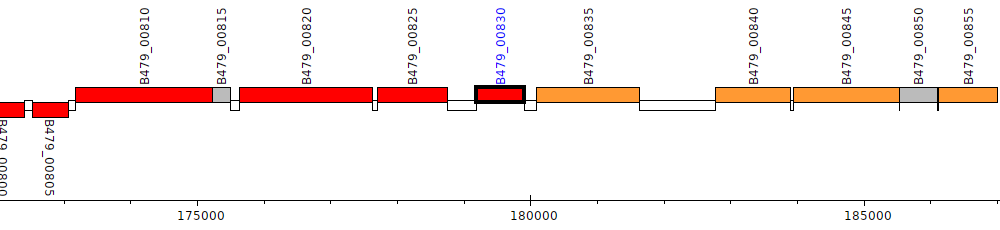
Pseudomonas putida HB3267, B479_00830
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00947
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004089 | carbonate dehydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00947
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.1050.10 | Carbonic anhydrase | IPR036874 | Carbonic anhydrase superfamily | 8 | 223 | 3.7E-70 |
| Pfam | PF00484 | Carbonic anhydrase | IPR001765 | Carbonic anhydrase | 55 | 210 | 1.2E-51 |
| CDD | cd00884 | beta_CA_cladeB | IPR045066 | Beta carbonic anhydrases, cladeB | 28 | 214 | 1.14136E-103 |
| SUPERFAMILY | SSF53056 | beta-carbonic anhydrase, cab | IPR036874 | Carbonic anhydrase superfamily | 16 | 224 | 3.92E-69 |
| PANTHER | PTHR11002 | CARBONIC ANHYDRASE | IPR001765 | Carbonic anhydrase | 17 | 216 | 1.1E-64 |
| FunFam | G3DSA:3.40.1050.10:FF:000003 | Carbonic anhydrase | - | - | 17 | 223 | 1.8E-75 |
| SMART | SM00947 | Pro_CA_2 | IPR001765 | Carbonic anhydrase | 48 | 215 | 8.4E-66 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.