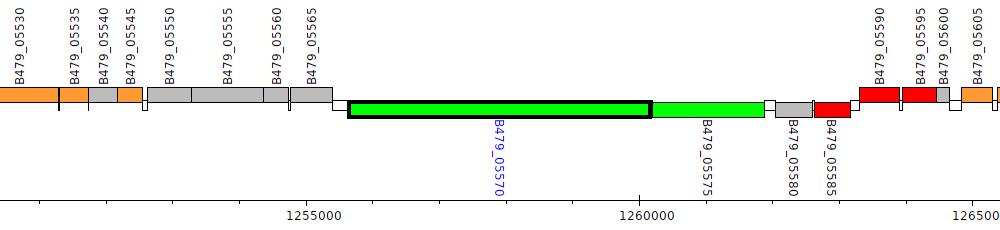
Pseudomonas putida HB3267, B479_05570
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh03070 | Bacterial secretion system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1066 | 1094 | - |
| Pfam | PF05860 | TPS secretion domain | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 43 | 291 | 3.0E-38 |
| NCBIfam | TIGR01901 | JCVI: filamentous hemagglutinin family N-terminal domain | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 68 | 158 | 3.1E-14 |
| Pfam | PF13332 | Hemagglutinin repeat | IPR025157 | Hemagglutinin repeat | 306 | 428 | 4.2E-5 |
| SMART | SM00912 | Haemagg_act_2a | IPR008638 | Filamentous haemagglutinin FhaB/tRNA nuclease CdiA-like, TPS domain | 50 | 170 | 5.6E-41 |
| Pfam | PF13332 | Hemagglutinin repeat | IPR025157 | Hemagglutinin repeat | 1024 | 1157 | 1.0E-7 |
| Gene3D | G3DSA:2.160.20.10 | - | IPR012334 | Pectin lyase fold | 41 | 348 | 8.7E-74 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 987 | 1007 | - |
| Pfam | PF13332 | Hemagglutinin repeat | IPR025157 | Hemagglutinin repeat | 594 | 756 | 3.2E-7 |
| Pfam | PF13332 | Hemagglutinin repeat | IPR025157 | Hemagglutinin repeat | 1175 | 1330 | 1.4E-15 |
| Pfam | PF13332 | Hemagglutinin repeat | IPR025157 | Hemagglutinin repeat | 521 | 586 | 0.0046 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 654 | 685 | - |
| SUPERFAMILY | SSF51126 | Pectin lyase-like | IPR011050 | Pectin lyase fold/virulence factor | 46 | 294 | 5.02E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.