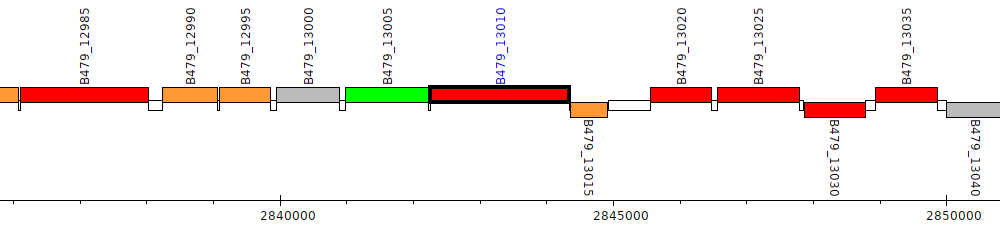
Pseudomonas putida HB3267, B479_13010
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006631 | fatty acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02737
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0070403 | NAD+ binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02737
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00725
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | fermentation to 2-methylbutanoate | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | docosahexaenoate biosynthesis III (6-desaturase, mammals) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 3-hydroxypropanoate/4-hydroxybutanate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-hydroxybenzoate biosynthesis III (plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to butanol I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-coumarate degradation (aerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh01212 | Fatty acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | fatty acid β-oxidation II (peroxisome) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | cholesterol degradation to androstenedione II (cholesterol dehydrogenase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to hexanol (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | unsaturated, even numbered fatty acid β-oxidation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | androstenedione degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh00362 | Benzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh00650 | Butanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | (4Z,7Z,10Z,13Z,16Z)-docosapentaenoate biosynthesis (6-desaturase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glutaryl-CoA degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | methyl <i>tert</i>-butyl ether degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | fatty acid salvage | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to butanol II (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | jasmonic acid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | <i>Spodoptera littoralis</i> pheromone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | (8<i>E</i>,10<i>E</i>)-dodeca-8,10-dienol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 4-coumarate degradation (anaerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | methyl ketone biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | benzoyl-CoA degradation I (aerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | cholesterol degradation to androstenedione I (cholesterol oxidase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 2-methylpropene degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | crotonate fermentation (to acetate and cyclohexane carboxylate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | (R)- and (S)-3-hydroxybutanoate biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuh00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF02737 | 3-hydroxyacyl-CoA dehydrogenase, NAD binding domain | IPR006176 | 3-hydroxyacyl-CoA dehydrogenase, NAD binding | 293 | 470 | 9.1E-58 |
| PANTHER | PTHR23309 | 3-HYDROXYACYL-COA DEHYROGENASE | - | - | 11 | 692 | 0.0 |
| FunFam | G3DSA:3.40.50.720:FF:000009 | Fatty oxidation complex, alpha subunit | - | - | 291 | 476 | 2.3E-60 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 599 | 686 | 6.38E-20 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 280 | 468 | 2.3E-64 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 474 | 616 | 2.38E-30 |
| SUPERFAMILY | SSF52096 | ClpP/crotonase | IPR029045 | ClpP/crotonase-like domain superfamily | 4 | 276 | 3.2E-61 |
| Gene3D | G3DSA:1.10.1040.50 | - | - | - | 476 | 693 | 1.8E-72 |
| Pfam | PF00725 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal domain | IPR006108 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal | 474 | 567 | 6.1E-25 |
| Pfam | PF00378 | Enoyl-CoA hydratase/isomerase | IPR001753 | Enoyl-CoA hydratase/isomerase | 10 | 193 | 1.2E-37 |
| Pfam | PF00725 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal domain | IPR006108 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal | 603 | 686 | 5.3E-8 |
| FunFam | G3DSA:1.10.1040.50:FF:000006 | Peroxisomal bifunctional enzyme | - | - | 476 | 694 | 2.6E-70 |
| CDD | cd06558 | crotonase-like | - | - | 5 | 189 | 5.84359E-56 |
| Gene3D | G3DSA:3.90.226.10 | - | - | - | 6 | 277 | 2.6E-78 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 291 | 472 | 3.52E-53 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.