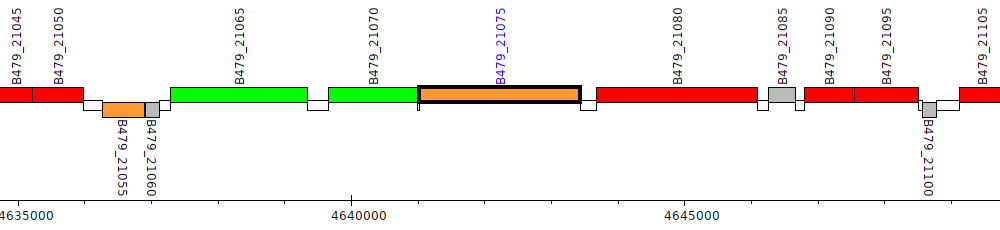
Pseudomonas putida HB3267, B479_21075
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0048038 | quinone binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd10280
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:cd10280
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016614 | oxidoreductase activity, acting on CH-OH group of donors |
Inferred from Sequence Model
Term mapped from: InterPro:cd10280
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00564 | ire1_9 | IPR018391 | Pyrrolo-quinoline quinone beta-propeller repeat | 358 | 399 | 13.0 |
| SMART | SM00564 | ire1_9 | IPR018391 | Pyrrolo-quinoline quinone beta-propeller repeat | 301 | 333 | 200.0 |
| NCBIfam | TIGR03074 | JCVI: membrane-bound PQQ-dependent dehydrogenase, glucose/quinate/shikimate family | IPR017511 | PQQ-dependent membrane bound dehydrogenase | 16 | 801 | 0.0 |
| CDD | cd10280 | PQQ_mGDH | IPR017511 | PQQ-dependent membrane bound dehydrogenase | 171 | 801 | 0.0 |
| Pfam | PF13360 | PQQ-like domain | IPR002372 | Pyrrolo-quinoline quinone repeat | 219 | 398 | 1.7E-5 |
| Gene3D | G3DSA:2.140.10.10 | - | - | - | 152 | 802 | 0.0 |
| SMART | SM00564 | ire1_9 | IPR018391 | Pyrrolo-quinoline quinone beta-propeller repeat | 223 | 254 | 1.1E-5 |
| Pfam | PF01011 | PQQ enzyme repeat | IPR002372 | Pyrrolo-quinoline quinone repeat | 736 | 766 | 6.8E-6 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 531 | 552 | - |
| SMART | SM00564 | ire1_9 | IPR018391 | Pyrrolo-quinoline quinone beta-propeller repeat | 426 | 477 | 88.0 |
| SUPERFAMILY | SSF50998 | Quinoprotein alcohol dehydrogenase-like | IPR011047 | Quinoprotein alcohol dehydrogenase-like superfamily | 165 | 801 | 0.0 |
| PANTHER | PTHR32303 | QUINOPROTEIN ALCOHOL DEHYDROGENASE (CYTOCHROME C) | - | - | 164 | 799 | 2.0E-61 |
| Pfam | PF01011 | PQQ enzyme repeat | IPR002372 | Pyrrolo-quinoline quinone repeat | 675 | 710 | 7.6E-6 |
| SMART | SM00564 | ire1_9 | IPR018391 | Pyrrolo-quinoline quinone beta-propeller repeat | 724 | 757 | 0.0022 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.