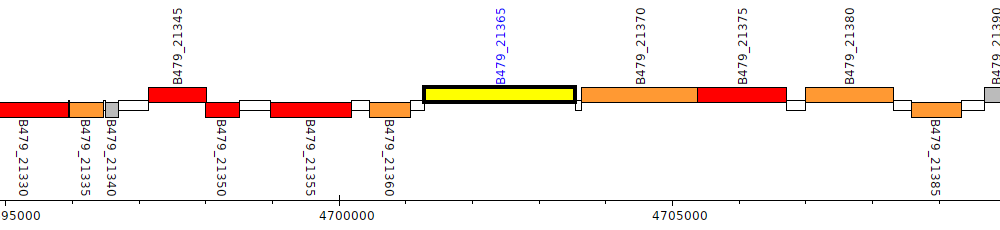
Pseudomonas putida HB3267, B479_21365
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00933
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:PF00933
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh00500 | Starch and sucrose metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuh00460 | Cyanoamino acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.20.20.300 | - | IPR036962 | Glycoside hydrolase, family 3, N-terminal domain superfamily | 17 | 359 | 2.2E-107 |
| PANTHER | PTHR30620 | PERIPLASMIC BETA-GLUCOSIDASE-RELATED | - | - | 18 | 636 | 0.0 |
| Pfam | PF00933 | Glycosyl hydrolase family 3 N terminal domain | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 35 | 335 | 1.7E-83 |
| PRINTS | PR00133 | Glycosyl hydrolase family 3 signature | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 188 | 204 | 3.1E-24 |
| PRINTS | PR00133 | Glycosyl hydrolase family 3 signature | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 151 | 167 | 3.1E-24 |
| FunFam | G3DSA:2.60.40.10:FF:000495 | Periplasmic beta-glucosidase | - | - | 638 | 753 | 3.2E-39 |
| FunFam | G3DSA:3.40.50.1700:FF:000004 | Periplasmic beta-glucosidase | - | - | 366 | 636 | 4.6E-121 |
| SMART | SM01217 | Fn3_like_2 | IPR026891 | Fibronectin type III-like domain | 673 | 742 | 2.6E-29 |
| PRINTS | PR00133 | Glycosyl hydrolase family 3 signature | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 86 | 102 | 3.1E-24 |
| PRINTS | PR00133 | Glycosyl hydrolase family 3 signature | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 258 | 276 | 3.1E-24 |
| Gene3D | G3DSA:2.60.40.10 | Immunoglobulins | IPR013783 | Immunoglobulin-like fold | 637 | 752 | 5.7E-34 |
| SUPERFAMILY | SSF52279 | Beta-D-glucan exohydrolase, C-terminal domain | IPR036881 | Glycoside hydrolase family 3 C-terminal domain superfamily | 380 | 639 | 5.49E-58 |
| FunFam | G3DSA:3.20.20.300:FF:000005 | Periplasmic beta-glucosidase | - | - | 16 | 357 | 0.0 |
| Pfam | PF14310 | Fibronectin type III-like domain | IPR026891 | Fibronectin type III-like domain | 674 | 742 | 3.8E-21 |
| SUPERFAMILY | SSF51445 | (Trans)glycosidases | IPR017853 | Glycoside hydrolase superfamily | 18 | 379 | 5.9E-99 |
| Gene3D | G3DSA:3.40.50.1700 | - | IPR036881 | Glycoside hydrolase family 3 C-terminal domain superfamily | 366 | 636 | 4.4E-88 |
| Pfam | PF01915 | Glycosyl hydrolase family 3 C-terminal domain | IPR002772 | Glycoside hydrolase family 3 C-terminal domain | 380 | 636 | 1.2E-60 |
| PRINTS | PR00133 | Glycosyl hydrolase family 3 signature | IPR001764 | Glycoside hydrolase, family 3, N-terminal | 105 | 124 | 3.1E-24 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.