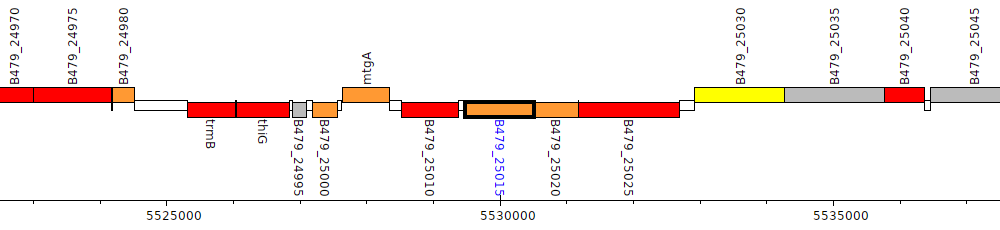
Pseudomonas putida HB3267, B479_25015
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0051301 | cell division |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF003097
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF003097
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuh02010 | ABC transporters | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.30.70.3040:FF:000002 | Cell division protein FtsX | - | - | 86 | 194 | 4.4E-55 |
| Pfam | PF18075 | FtsX extracellular domain | IPR040690 | FtsX, extracellular domain | 100 | 190 | 1.1E-16 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 32 | - |
| Gene3D | G3DSA:3.30.70.3040 | - | - | - | 86 | 194 | 2.1E-15 |
| Pfam | PF02687 | FtsX-like permease family | IPR003838 | ABC3 transporter permease protein domain | 216 | 331 | 5.1E-7 |
| PANTHER | PTHR47755 | CELL DIVISION PROTEIN FTSX | IPR004513 | Cell division protein FtsX | 40 | 335 | 5.1E-86 |
| PIRSF | PIRSF003097 | FtsX | IPR004513 | Cell division protein FtsX | 42 | 336 | 4.1E-66 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 18 | 32 | - |
| NCBIfam | TIGR00439 | JCVI: permease-like cell division protein FtsX | IPR047590 | Cell division protein FtsX, proteobacteria | 46 | 335 | 1.7E-101 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.