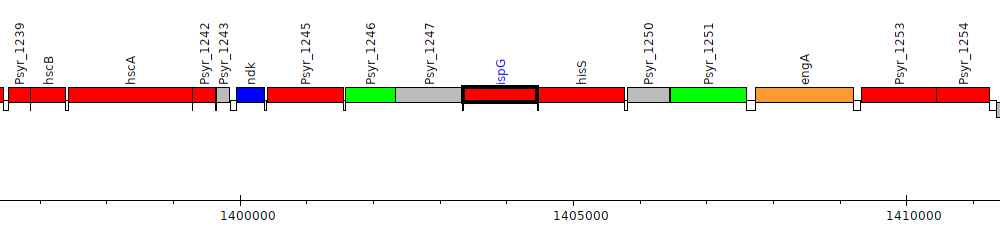
Pseudomonas syringae pv. syringae B728a, Psyr_1248 (ispG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005506 | iron ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF004640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016114 | terpenoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00159
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0044237 | cellular metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.20.20.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046429 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00159
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008299 | isoprenoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF004640
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psb01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb00900 | Terpenoid backbone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.20.20.20:FF:000001 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase (flavodoxin) | - | - | 1 | 261 | 3.0E-129 |
| PIRSF | PIRSF004640 | IspG | IPR016425 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase, bacterial | 1 | 362 | 0.0 |
| Pfam | PF04551 | GcpE protein | IPR004588 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase, bacterial-type | 13 | 352 | 0.0 |
| NCBIfam | TIGR00612 | JCVI: (E)-4-hydroxy-3-methylbut-2-enyl-diphosphate synthase | IPR004588 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase, bacterial-type | 9 | 351 | 4.1E-130 |
| Gene3D | G3DSA:3.20.20.20 | - | IPR011005 | Dihydropteroate synthase-like | 1 | 261 | 4.6E-107 |
| SUPERFAMILY | SSF51412 | Inosine monophosphate dehydrogenase (IMPDH) | - | - | 24 | 132 | 8.06E-10 |
| Gene3D | G3DSA:3.30.413.10 | Sulfite Reductase Hemoprotein, domain 1 | IPR045854 | Nitrite and sulphite reductase 4Fe-4S domain-like superfamily | 262 | 361 | 1.3E-26 |
| SUPERFAMILY | SSF56014 | Nitrite and sulphite reductase 4Fe-4S domain-like | IPR045854 | Nitrite and sulphite reductase 4Fe-4S domain-like superfamily | 265 | 352 | 5.23E-12 |
| Hamap | MF_00159 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase (ferredoxin) [ispG]. | IPR004588 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase, bacterial-type | 7 | 359 | 35.355534 |
| PANTHER | PTHR30454 | 4-HYDROXY-3-METHYLBUT-2-EN-1-YL DIPHOSPHATE SYNTHASE | IPR004588 | 4-hydroxy-3-methylbut-2-en-1-yl diphosphate synthase, bacterial-type | 7 | 252 | 0.0 |
| Coils | Coil | Coil | - | - | 340 | 364 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.