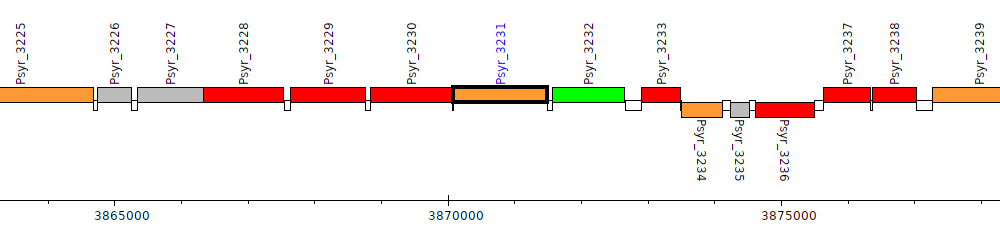
Pseudomonas syringae pv. syringae B728a, Psyr_3231
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | xanthan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | acetan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR30576 | COLANIC BIOSYNTHESIS UDP-GLUCOSE LIPID CARRIER TRANSFERASE | - | - | 131 | 469 | 6.6E-87 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 141 | 268 | 4.8E-13 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 142 | 254 | 3.69E-9 |
| NCBIfam | TIGR03025 | JCVI: exopolysaccharide biosynthesis polyprenyl glycosylphosphotransferase | IPR017475 | Exopolysaccharide biosynthesis polyprenyl glycosylphosphotransferase | 23 | 468 | 0.0 |
| Pfam | PF02397 | Bacterial sugar transferase | IPR003362 | Bacterial sugar transferase | 279 | 461 | 3.8E-65 |
| NCBIfam | TIGR03023 | JCVI: undecaprenyl-phosphate glucose phosphotransferase | IPR017473 | Undecaprenyl-phosphate glucose phosphotransferase, WcaJ | 28 | 469 | 0.0 |
| Pfam | PF13727 | CoA-binding domain | - | - | 66 | 243 | 3.8E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.