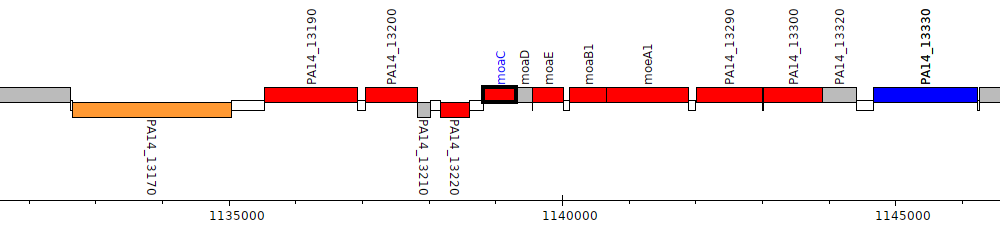
Pseudomonas aeruginosa UCBPP-PA14, PA14_13230 (moaC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006777 | Mo-molybdopterin cofactor biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00581
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau04122 | Sulfur relay system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00790 | Folate biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.30.70.640:FF:000001 | Cyclic pyranopterin monophosphate synthase | - | - | 1 | 158 | 7.2E-74 |
| Pfam | PF01967 | MoaC family | IPR002820 | Molybdopterin cofactor biosynthesis C (MoaC) domain | 13 | 147 | 2.4E-57 |
| Gene3D | G3DSA:3.30.70.640 | Molybdopterin cofactor biosynthesis C (MoaC) domain | IPR036522 | Molybdopterin cofactor biosynthesis C (MoaC) domain superfamily | 1 | 158 | 1.1E-68 |
| NCBIfam | TIGR00581 | JCVI: cyclic pyranopterin monophosphate synthase MoaC | IPR023045 | Molybdenum cofactor biosynthesis C | 2 | 149 | 2.2E-69 |
| CDD | cd01420 | MoaC_PE | IPR047594 | Molybdenum cofactor biosynthesis C, bacteria/eukaryotes | 13 | 151 | 1.13436E-82 |
| Hamap | MF_01224_B | Cyclic pyranopterin monophosphate synthase [moaC]. | IPR047594 | Molybdenum cofactor biosynthesis C, bacteria/eukaryotes | 1 | 155 | 35.503487 |
| SUPERFAMILY | SSF55040 | Molybdenum cofactor biosynthesis protein C, MoaC | IPR036522 | Molybdopterin cofactor biosynthesis C (MoaC) domain superfamily | 9 | 152 | 3.79E-60 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.