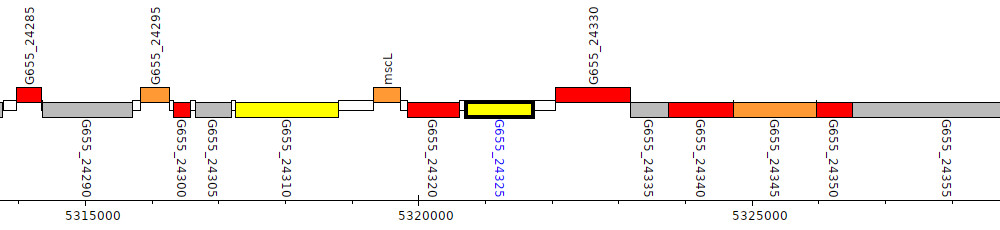
Pseudomonas aeruginosa B136-33, G655_24325
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0030288 | outer membrane-bounded periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006470
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psg02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.40.190.170:FF:000001 | TRAP dicarboxylate transporter, DctP subunit | - | - | 27 | 329 | 8.6E-111 |
| PANTHER | PTHR33376 | - | IPR018389 | TRAP transporter solute receptor DctP | 1 | 328 | 6.8E-106 |
| Gene3D | G3DSA:3.40.190.170 | - | IPR038404 | TRAP transporter solute receptor DctP superfamily | 28 | 331 | 3.2E-82 |
| NCBIfam | TIGR00787 | JCVI: DctP family TRAP transporter solute-binding subunit | IPR004682 | TRAP transporter solute receptor, DctP family | 32 | 277 | 5.8E-65 |
| NCBIfam | NF037995 | NCBIFAM: TRAP transporter substrate-binding protein DctP | - | - | 31 | 302 | 7.8E-46 |
| Pfam | PF03480 | Bacterial extracellular solute-binding protein, family 7 | IPR018389 | TRAP transporter solute receptor DctP | 32 | 314 | 9.3E-97 |
| PIRSF | PIRSF006470 | DctB | IPR004682 | TRAP transporter solute receptor, DctP family | 1 | 331 | 1.2E-117 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.