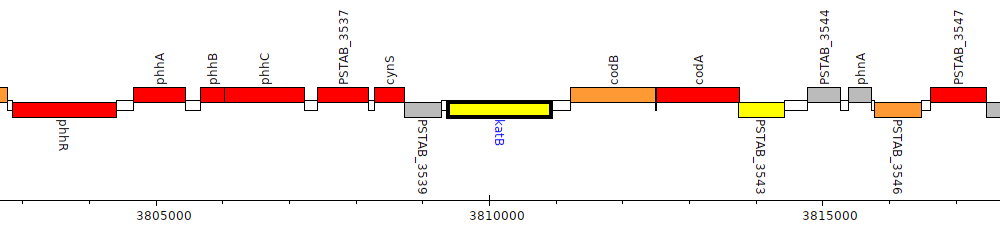
Pseudomonas stutzeri ATCC 17588 = LMG 11199, PSTAB_3540 (katB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004096 | catalase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038928
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006979 | response to oxidative stress |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038928
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0020037 | heme binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF038928
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psz00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz00380 | Tryptophan metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psz01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 46 | 69 | 4.8E-67 |
| Pfam | PF00199 | Catalase | IPR011614 | Catalase core domain | 33 | 409 | 0.0 |
| Gene3D | G3DSA:2.40.180.10 | Catalase core domain | - | - | 30 | 512 | 0.0 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 381 | 406 | - |
| PANTHER | PTHR11465 | CATALASE | IPR018028 | Catalase, mono-functional, haem-containing | 31 | 495 | 0.0 |
| Pfam | PF06628 | Catalase-related immune-responsive | IPR010582 | Catalase immune-responsive domain | 434 | 494 | 3.4E-15 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 108 | 126 | 4.8E-67 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 37 | 55 | - |
| SMART | SM01060 | Catalase_2 | IPR011614 | Catalase core domain | 33 | 413 | 0.0 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 311 | 338 | 4.8E-67 |
| FunFam | G3DSA:2.40.180.10:FF:000002 | Catalase | - | - | 29 | 502 | 0.0 |
| CDD | cd08154 | catalase_clade_1 | - | - | 32 | 496 | 0.0 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 343 | 369 | 4.8E-67 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 148 | 166 | 4.8E-67 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 380 | 416 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 34 | 55 | - |
| PIRSF | PIRSF038928 | Catalase_clade1-3 | IPR024711 | Catalase, mono-functional, haem-containing, clades 1 and 3 | 24 | 502 | 0.0 |
| PRINTS | PR00067 | Catalase signature | IPR018028 | Catalase, mono-functional, haem-containing | 129 | 146 | 4.8E-67 |
| SUPERFAMILY | SSF56634 | Heme-dependent catalase-like | IPR020835 | Catalase superfamily | 28 | 501 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.