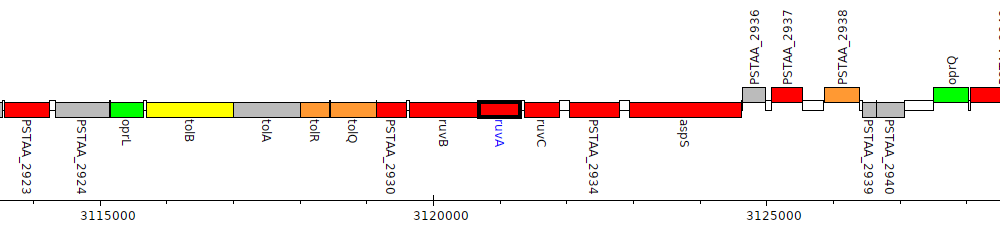
Pseudomonas stutzeri DSM 4166, PSTAA_2932 (ruvA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00031
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003678 | DNA helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00031
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009379 | Holliday junction helicase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PF07499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00278
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009378 | four-way junction helicase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006310 | DNA recombination |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00031
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF07499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psr03440 | Homologous recombination | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14520 | Helix-hairpin-helix domain | - | - | 72 | 130 | 3.1E-17 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 1 | 63 | 3.94E-19 |
| Gene3D | G3DSA:1.10.8.10 | - | - | - | 155 | 201 | 3.1E-19 |
| Pfam | PF01330 | RuvA N terminal domain | IPR013849 | DNA helicase, Holliday junction RuvA type, domain I, bacterial | 1 | 62 | 9.7E-24 |
| SMART | SM00278 | HhH1_4 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 108 | 127 | 0.086 |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 1 | 66 | 3.6E-28 |
| Hamap | MF_00031 | Holliday junction branch migration complex subunit RuvA [ruvA]. | IPR000085 | Holliday junction branch migration complex subunit RuvA | 1 | 201 | 32.266731 |
| CDD | cd14332 | UBA_RuvA_C | IPR011114 | Holliday junction DNA helicase RuvA, C-terminal | 156 | 198 | 1.47497E-11 |
| Coils | Coil | Coil | - | - | 118 | 138 | - |
| SUPERFAMILY | SSF46929 | DNA helicase RuvA subunit, C-terminal domain | IPR036267 | RuvA, C-terminal domain superfamily | 157 | 200 | 6.54E-11 |
| Gene3D | G3DSA:1.10.150.20 | - | - | - | 67 | 140 | 1.6E-28 |
| Pfam | PF07499 | RuvA, C-terminal domain | IPR011114 | Holliday junction DNA helicase RuvA, C-terminal | 157 | 200 | 2.2E-15 |
| NCBIfam | TIGR00084 | JCVI: Holliday junction branch migration protein RuvA | IPR000085 | Holliday junction branch migration complex subunit RuvA | 1 | 199 | 2.2E-61 |
| SUPERFAMILY | SSF47781 | RuvA domain 2-like | IPR010994 | RuvA domain 2-like | 66 | 136 | 5.0E-22 |
| SMART | SM00278 | HhH1_4 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 73 | 92 | 0.72 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.