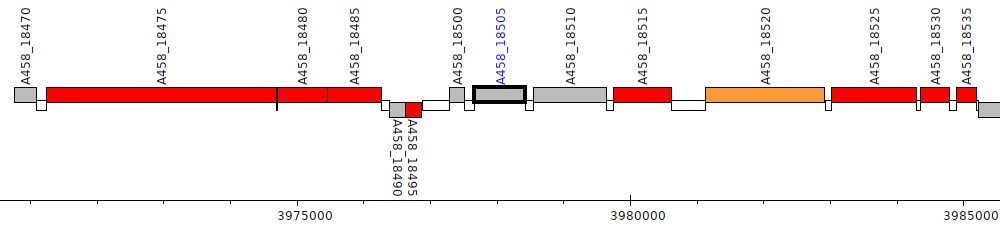
Pseudomonas stutzeri CCUG 29243, A458_18505
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006265 | DNA topological change |
Inferred from Sequence Model
Term mapped from: InterPro:PF01396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005694 | chromosome |
Inferred from Sequence Model
Term mapped from: InterPro:PF01396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003916 | DNA topoisomerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01396
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF57783 | Zinc beta-ribbon | - | - | 203 | 254 | 3.23E-7 |
| Pfam | PF08378 | Nuclease-related domain | IPR011528 | Nuclease-related domain, NERD | 33 | 144 | 2.5E-27 |
| Pfam | PF01396 | Topoisomerase DNA binding C4 zinc finger | IPR013498 | DNA topoisomerase, type IA, zn finger | 217 | 253 | 3.4E-9 |
| Gene3D | G3DSA:3.30.65.10 | Bacterial Topoisomerase I, domain 1 | - | - | 203 | 256 | 6.8E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.