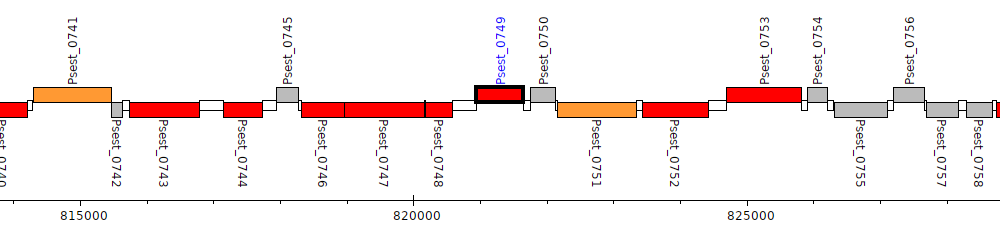
Pseudomonas stutzeri RCH2, Psest_0749
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0045892 | negative regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF08362
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR00455
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF08362 | YcdC-like protein, C-terminal region | IPR013573 | Transcription regulator YcdC, C-terminal | 94 | 236 | 2.7E-57 |
| PRINTS | PR00455 | TetR bacterial regulatory protein HTH signature | IPR001647 | DNA-binding HTH domain, TetR-type | 68 | 91 | 5.0E-8 |
| SUPERFAMILY | SSF48498 | Tetracyclin repressor-like, C-terminal domain | IPR036271 | Tetracyclin repressor-like, C-terminal domain superfamily | 111 | 236 | 3.42E-42 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 28 | - |
| Pfam | PF00440 | Bacterial regulatory proteins, tetR family | IPR001647 | DNA-binding HTH domain, TetR-type | 47 | 91 | 5.5E-14 |
| PRINTS | PR00455 | TetR bacterial regulatory protein HTH signature | IPR001647 | DNA-binding HTH domain, TetR-type | 47 | 60 | 5.0E-8 |
| SUPERFAMILY | SSF46689 | Homeodomain-like | IPR009057 | Homeobox-like domain superfamily | 36 | 115 | 3.75E-19 |
| Gene3D | G3DSA:1.10.357.10 | Tetracycline Repressor, domain 2 | - | - | 94 | 236 | 4.8E-51 |
| PANTHER | PTHR30328 | TRANSCRIPTIONAL REPRESSOR | - | - | 37 | 235 | 1.7E-22 |
| Gene3D | G3DSA:1.10.10.60 | - | - | - | 28 | 93 | 2.4E-22 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.