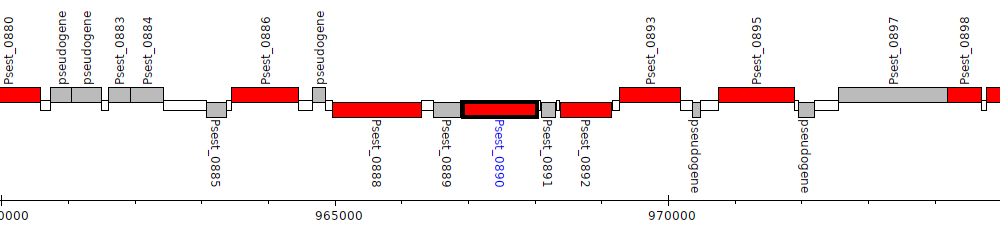
Pseudomonas stutzeri RCH2, Psest_0890
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051903 | S-(hydroxymethyl)glutathione dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd08300
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006069 | ethanol oxidation |
Inferred from Sequence Model
Term mapped from: InterPro:cd08300
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd08300
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psh01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | protein <i>S</i>-nitrosylation and denitrosylation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | chitin degradation to ethanol | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | (<i>S</i>)-propane-1,2-diol degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-leucine degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-isoleucine degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | salidroside biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to isobutanol (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psh00350 | Tyrosine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psh01220 | Degradation of aromatic compounds | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 3-methylbutanol biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh00010 | Glycolysis / Gluconeogenesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | acetaldehyde biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to ethanol II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-valine degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to ethanol III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh00626 | Naphthalene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | pyruvate fermentation to ethanol I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | serotonin degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-methionine degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | noradrenaline and adrenaline degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-tryptophan degradation V (side chain pathway) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-phenylalanine degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh00680 | Methane metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | formaldehyde oxidation II (glutathione-dependent) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | butanol and isobutanol biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | psh00625 | Chloroalkane and chloroalkene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | phenylethanol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF50129 | GroES-like | IPR011032 | GroES-like superfamily | 318 | 369 | 1.05E-11 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 161 | 333 | 5.16E-46 |
| NCBIfam | TIGR02818 | JCVI: S-(hydroxymethyl)glutathione dehydrogenase/class III alcohol dehydrogenase | IPR014183 | Alcohol dehydrogenase class III | 2 | 369 | 0.0 |
| SUPERFAMILY | SSF50129 | GroES-like | IPR011032 | GroES-like superfamily | 1 | 185 | 1.28E-70 |
| Gene3D | G3DSA:3.90.180.10 | - | - | - | 6 | 364 | 0.0 |
| FunFam | G3DSA:3.40.50.720:FF:000003 | S-(hydroxymethyl)glutathione dehydrogenase | - | - | 174 | 312 | 9.5E-66 |
| PANTHER | PTHR43880 | ALCOHOL DEHYDROGENASE | - | - | 2 | 367 | 0.0 |
| Pfam | PF08240 | Alcohol dehydrogenase GroES-like domain | IPR013154 | Alcohol dehydrogenase-like, N-terminal | 28 | 154 | 5.7E-26 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 174 | 312 | 0.0 |
| SMART | SM00829 | PKS_ER_names_mod | IPR020843 | Polyketide synthase, enoylreductase domain | 10 | 367 | 1.2E-4 |
| Pfam | PF00107 | Zinc-binding dehydrogenase | IPR013149 | Alcohol dehydrogenase-like, C-terminal | 197 | 317 | 5.8E-23 |
| CDD | cd08300 | alcohol_DH_class_III | IPR014183 | Alcohol dehydrogenase class III | 1 | 368 | 0.0 |
| FunFam | G3DSA:3.90.180.10:FF:000001 | S-(hydroxymethyl)glutathione dehydrogenase | - | - | 6 | 188 | 9.3E-87 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.