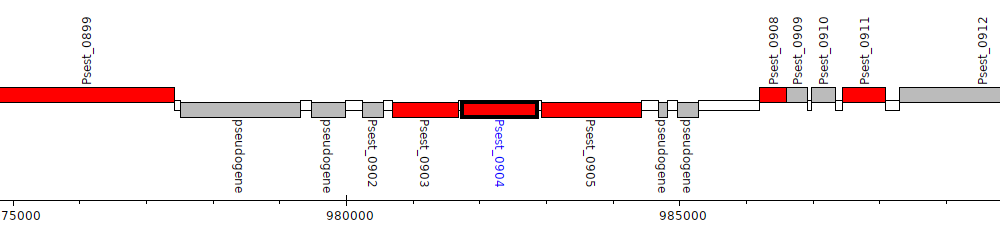
Pseudomonas stutzeri RCH2, Psest_0904
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psh00633 | Nitrotoluene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psh01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR22893 | NADH OXIDOREDUCTASE-RELATED | IPR045247 | Oxidoreductase Oye-like | 4 | 371 | 8.0E-113 |
| CDD | cd02933 | OYE_like_FMN | - | - | 5 | 355 | 0.0 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 3 | 374 | 0.0 |
| Pfam | PF00724 | NADH:flavin oxidoreductase / NADH oxidase family | IPR001155 | NADH:flavin oxidoreductase/NADH oxidase, N-terminal | 5 | 351 | 7.2E-83 |
| SUPERFAMILY | SSF51395 | FMN-linked oxidoreductases | - | - | 1 | 373 | 4.89E-123 |
| FunFam | G3DSA:3.20.20.70:FF:000059 | N-ethylmaleimide reductase, FMN-linked | - | - | 3 | 374 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.