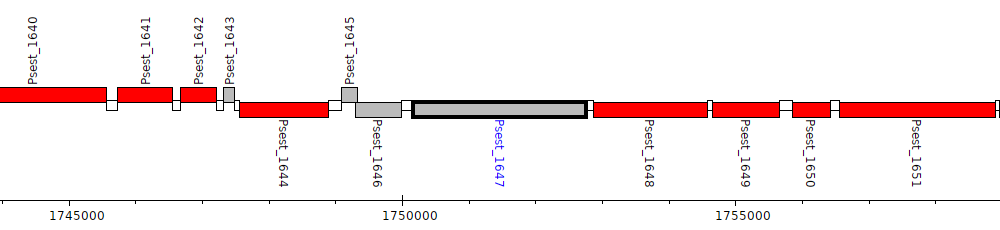
Pseudomonas stutzeri RCH2, Psest_1647
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF08494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016818 | hydrolase activity, acting on acid anhydrides, in phosphorus-containing anhydrides |
Inferred from Sequence Model
Term mapped from: InterPro:PF08494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00270
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 9 | 208 | 7.8E-38 |
| Pfam | PF08494 | DEAD/H associated | IPR013701 | DEAD/H associated | 602 | 788 | 3.3E-51 |
| Coils | Coil | Coil | - | - | 747 | 774 | - |
| SMART | SM00487 | ultradead3 | IPR014001 | Helicase superfamily 1/2, ATP-binding domain | 16 | 236 | 8.9E-23 |
| CDD | cd18796 | SF2_C_LHR | - | - | 221 | 367 | 4.74814E-64 |
| NCBIfam | TIGR04121 | JCVI: ligase-associated DNA damage response DEXH box helicase | IPR026362 | DEXH box helicase, DNA ligase-associated | 10 | 817 | 0.0 |
| SMART | SM00490 | helicmild6 | IPR001650 | Helicase, C-terminal | 272 | 353 | 2.1E-5 |
| Pfam | PF00271 | Helicase conserved C-terminal domain | IPR001650 | Helicase, C-terminal | 252 | 354 | 1.4E-10 |
| Pfam | PF19306 | Helicase Lhr winged helix domain | IPR045628 | Helicase Lhr-like, winged helix domain | 396 | 544 | 6.3E-59 |
| Pfam | PF00270 | DEAD/DEAH box helicase | IPR011545 | DEAD/DEAH box helicase domain | 22 | 199 | 7.7E-22 |
| PANTHER | PTHR47962 | ATP-DEPENDENT HELICASE LHR-RELATED-RELATED | - | - | 10 | 804 | 0.0 |
| PIRSF | PIRSF037307 | Lhr-like_helic | IPR017170 | Lhr-like ATP-dependent RNA helicase, predicted | 2 | 819 | 0.0 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 27 | 365 | 2.05E-41 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 817 | 863 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 836 | 857 | - |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 244 | 397 | 8.1E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.