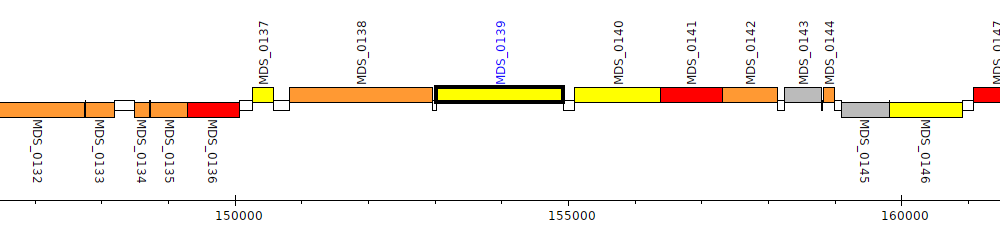
Pseudomonas mendocina NK-01, MDS_0139
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050304 | nitrous-oxide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04244
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005509 | calcium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04244
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.130.10.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005507 | copper ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR04244
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pmk01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | nitrate reduction VII (denitrification) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pmk00910 | Nitrogen metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | nitrifier denitrification | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF18793 | Nitrous oxide reductase propeller repeat 2 | IPR041142 | Nitrous oxide reductase, propeller repeat 2 | 167 | 237 | 1.7E-32 |
| SUPERFAMILY | SSF50974 | Nitrous oxide reductase, N-terminal domain | IPR011045 | Nitrous oxide reductase, N-terminal | 57 | 502 | 1.96E-128 |
| SUPERFAMILY | SSF49503 | Cupredoxins | IPR008972 | Cupredoxin | 505 | 633 | 9.64E-37 |
| Gene3D | G3DSA:2.60.40.420 | - | IPR008972 | Cupredoxin | 515 | 632 | 7.1E-38 |
| FunFam | G3DSA:2.130.10.10:FF:001102 | Nitrous-oxide reductase | - | - | 54 | 513 | 0.0 |
| Pfam | PF18764 | Nitrous oxide reductase propeller repeat | IPR041114 | Nitrous oxide reductase, propeller repeat 1 | 420 | 490 | 4.3E-35 |
| Gene3D | G3DSA:2.130.10.10 | - | IPR015943 | WD40/YVTN repeat-like-containing domain superfamily | 54 | 513 | 0.0 |
| CDD | cd04223 | N2OR_C | IPR034205 | Nitrous-oxide reductase, C-terminal | 539 | 633 | 1.57913E-52 |
| NCBIfam | TIGR01409 | JCVI: twin-arginine translocation signal domain | IPR019546 | Twin-arginine translocation pathway, signal sequence, bacterial/archaeal | 14 | 45 | 0.0026 |
| PANTHER | PTHR42838 | CYTOCHROME C OXIDASE SUBUNIT II | - | - | 3 | 633 | 0.0 |
| Hamap | MF_00716 | Nitrous-oxide reductase [nosZ]. | IPR023644 | Nitrous-oxide reductase | 2 | 633 | 62.883144 |
| NCBIfam | TIGR04244 | JCVI: TAT-dependent nitrous-oxide reductase | IPR023644 | Nitrous-oxide reductase | 14 | 632 | 0.0 |
| FunFam | G3DSA:2.60.40.420:FF:000095 | Nitrous-oxide reductase | - | - | 514 | 633 | 1.2E-74 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.