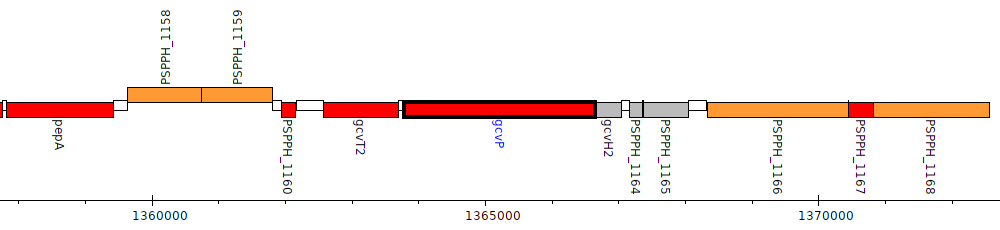
Pseudomonas savastanoi pv. phaseolicola 1448A, PSPPH_1162 (gcvP)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006544 | glycine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00461
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004375 | glycine dehydrogenase (decarboxylating) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00461
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006546 | glycine catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02347
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psp01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp00630 | Glyoxylate and dicarboxylate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psp00260 | Glycine, serine and threonine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00711 | Glycine dehydrogenase (decarboxylating) [gcvP]. | IPR003437 | Glycine dehydrogenase (decarboxylating) | 7 | 951 | 26.147501 |
| Pfam | PF02347 | Glycine cleavage system P-protein | IPR020581 | Glycine cleavage system P protein | 467 | 734 | 5.3E-10 |
| FunFam | G3DSA:3.40.640.10:FF:000007 | glycine dehydrogenase (Decarboxylating), mitochondrial | - | - | 522 | 753 | 1.1E-120 |
| Gene3D | G3DSA:3.90.1150.10 | Aspartate Aminotransferase, domain 1 | IPR015422 | Pyridoxal phosphate-dependent transferase, small domain | 83 | 410 | 6.6E-85 |
| FunFam | G3DSA:3.40.640.10:FF:000005 | Glycine dehydrogenase (decarboxylating), mitochondrial | - | - | 94 | 354 | 0.0 |
| NCBIfam | TIGR00461 | JCVI: aminomethyl-transferring glycine dehydrogenase | IPR003437 | Glycine dehydrogenase (decarboxylating) | 16 | 945 | 0.0 |
| Pfam | PF02347 | Glycine cleavage system P-protein | IPR020581 | Glycine cleavage system P protein | 16 | 440 | 0.0 |
| Gene3D | G3DSA:3.90.1150.10 | Aspartate Aminotransferase, domain 1 | IPR015422 | Pyridoxal phosphate-dependent transferase, small domain | 765 | 899 | 6.2E-46 |
| Gene3D | G3DSA:3.40.640.10 | - | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 94 | 354 | 6.6E-85 |
| FunFam | G3DSA:3.90.1150.10:FF:000007 | Glycine dehydrogenase (decarboxylating), mitochondrial | - | - | 761 | 899 | 1.5E-72 |
| Gene3D | G3DSA:3.40.640.10 | - | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 523 | 764 | 6.1E-72 |
| SUPERFAMILY | SSF53383 | PLP-dependent transferases | IPR015424 | Pyridoxal phosphate-dependent transferase | 16 | 441 | 4.96E-100 |
| CDD | cd00613 | GDC-P | IPR020581 | Glycine cleavage system P protein | 483 | 868 | 0.0 |
| PANTHER | PTHR11773 | GLYCINE DEHYDROGENASE, DECARBOXYLATING | IPR020581 | Glycine cleavage system P protein | 7 | 951 | 0.0 |
| SUPERFAMILY | SSF53383 | PLP-dependent transferases | IPR015424 | Pyridoxal phosphate-dependent transferase | 475 | 946 | 2.74E-103 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.