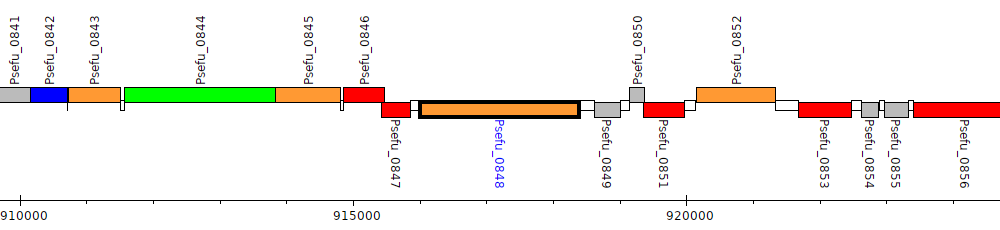
Pseudomonas fulva 12-X, Psefu_0848
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005215 | transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01525
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 448 | 739 | 5.9E-58 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 8 | 64 | 5.9E-13 |
| Gene3D | G3DSA:3.40.1110.10 | - | IPR023299 | P-type ATPase, cytoplasmic domain N | 492 | 614 | 5.9E-25 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 705 | 717 | 1.3E-19 |
| Pfam | PF00403 | Heavy-metal-associated domain | IPR006121 | Heavy metal-associated domain, HMA | 72 | 130 | 7.9E-12 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 479 | 782 | 8.73E-57 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 68 | 134 | 1.2E-20 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 464 | 479 | 7.8E-11 |
| SUPERFAMILY | SSF81665 | Calcium ATPase, transmembrane domain M | IPR023298 | P-type ATPase, transmembrane domain superfamily | 250 | 759 | 1.57E-15 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 201 | 220 | 7.8E-11 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 682 | 701 | 1.3E-19 |
| Gene3D | G3DSA:2.70.150.10 | - | - | - | 274 | 386 | 3.8E-30 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 481 | 495 | 1.3E-19 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 68 | 136 | 5.63E-19 |
| FunFam | G3DSA:3.30.70.100:FF:000005 | Copper-exporting P-type ATPase A | - | - | 1 | 67 | 1.4E-18 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 253 | 508 | 2.8E-41 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 630 | 640 | 1.3E-19 |
| Pfam | PF00122 | E1-E2 ATPase | - | - | 282 | 461 | 8.3E-53 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 3 | 67 | 6.6E-19 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 4 | 65 | 2.36E-16 |
| CDD | cd02094 | P-type_ATPase_Cu-like | - | - | 170 | 785 | 0.0 |
| NCBIfam | TIGR01525 | JCVI: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 216 | 784 | 0.0 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 84 | 98 | 7.8E-11 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 5 | 65 | 2.82155E-17 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 660 | 677 | 7.8E-11 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 404 | 418 | 7.8E-11 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 478 | 695 | 1.2E-40 |
| SFLD | SFLDF00027 | p-type atpase | IPR044492 | P-type ATPase, haloacid dehalogenase domain | 463 | 734 | 0.0 |
| FunFam | G3DSA:2.70.150.10:FF:000002 | Copper-transporting ATPase 1, putative | - | - | 269 | 385 | 1.2E-34 |
| SFLD | SFLDG00002 | C1.7: P-type atpase like | - | - | 463 | 734 | 0.0 |
| PANTHER | PTHR43520 | ATP7, ISOFORM B | - | - | 4 | 789 | 0.0 |
| PRINTS | PR00943 | Copper-transporting ATPase signature | - | - | 251 | 270 | 7.8E-11 |
| NCBIfam | TIGR01511 | JCVI: copper-translocating P-type ATPase | - | - | 198 | 785 | 0.0 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 71 | 133 | 1.7708E-18 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 601 | 761 | 5.3E-40 |
| SUPERFAMILY | SSF81653 | Calcium ATPase, transduction domain A | IPR008250 | P-type ATPase, A domain superfamily | 284 | 381 | 1.29E-22 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 331 | 345 | 1.3E-19 |
| FunFam | G3DSA:3.30.70.100:FF:000005 | Copper-exporting P-type ATPase A | - | - | 68 | 135 | 3.5E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.