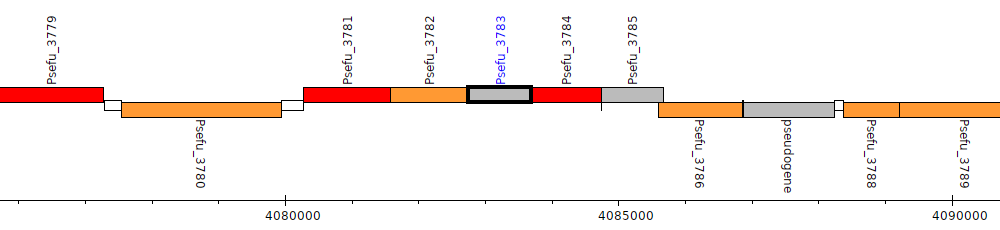
Pseudomonas fulva 12-X, Psefu_3783
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pfv02024 | Quorum sensing | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pfv02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00535 | Glycosyl transferase family 2 | IPR001173 | Glycosyltransferase 2-like | 50 | 133 | 9.6E-12 |
| SUPERFAMILY | SSF53448 | Nucleotide-diphospho-sugar transferases | IPR029044 | Nucleotide-diphospho-sugar transferases | 27 | 287 | 4.78E-33 |
| PANTHER | PTHR43179 | RHAMNOSYLTRANSFERASE WBBL | - | - | 47 | 285 | 2.7E-28 |
| Gene3D | G3DSA:3.90.550.10 | Spore Coat Polysaccharide Biosynthesis Protein SpsA; Chain A | IPR029044 | Nucleotide-diphospho-sugar transferases | 28 | 241 | 1.9E-24 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.