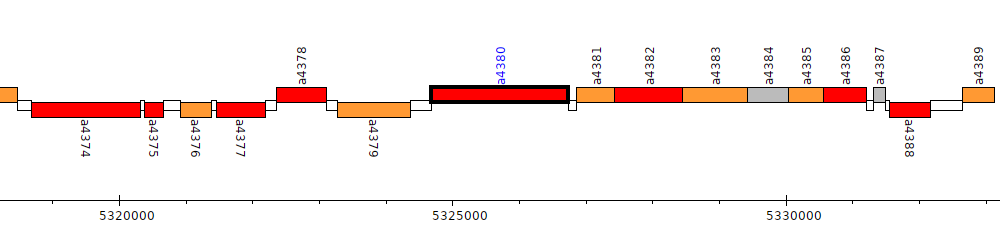
Pseudomonas brassicacearum subsp. brassicacearum NFM421, PSEBR_a4380
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006431 | methionyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR01041
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47323
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47323
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004825 | methionine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR01041
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00399
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000049 | tRNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01588
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pba00450 | Selenocompound metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pba00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:1.10.730.10:FF:000005 | Methionine--tRNA ligase | - | - | 388 | 526 | 2.5E-56 |
| NCBIfam | TIGR00399 | JCVI: methionine--tRNA ligase subunit beta | IPR004495 | Methionyl-tRNA synthetase, beta subunit, C-terminal | 558 | 681 | 2.9E-41 |
| Gene3D | G3DSA:3.40.50.620 | HUPs | IPR014729 | Rossmann-like alpha/beta/alpha sandwich fold | 7 | 383 | 0.0 |
| NCBIfam | TIGR00398 | JCVI: methionine--tRNA ligase | IPR014758 | Methionyl-tRNA synthetase | 7 | 535 | 0.0 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 250 | 261 | 2.6E-24 |
| CDD | cd02800 | tRNA_bind_EcMetRS_like | IPR004495 | Methionyl-tRNA synthetase, beta subunit, C-terminal | 575 | 681 | 1.81751E-46 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 290 | 305 | 2.6E-24 |
| SUPERFAMILY | SSF52374 | Nucleotidylyl transferase | - | - | 4 | 386 | 1.9E-98 |
| CDD | cd00814 | MetRS_core | IPR033911 | Methioninyl-tRNA synthetase core domain | 6 | 370 | 0.0 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 383 | 394 | 2.6E-24 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 41 | 55 | 2.6E-24 |
| SUPERFAMILY | SSF47323 | Anticodon-binding domain of a subclass of class I aminoacyl-tRNA synthetases | IPR009080 | Aminoacyl-tRNA synthetase, class Ia, anticodon-binding | 389 | 546 | 6.76E-40 |
| PANTHER | PTHR45765 | METHIONINE--TRNA LIGASE | IPR023458 | Methionine-tRNA ligase, type 1 | 3 | 674 | 0.0 |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 572 | 681 | 6.1E-35 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 9 | 22 | 2.6E-24 |
| Pfam | PF09334 | tRNA synthetases class I (M) | IPR015413 | Methionyl/Leucyl tRNA synthetase | 7 | 395 | 0.0 |
| Hamap | MF_00098 | Methionine--tRNA ligase [metG]. | IPR023458 | Methionine-tRNA ligase, type 1 | 5 | 545 | 39.309181 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 574 | 680 | 1.81E-28 |
| CDD | cd07957 | Anticodon_Ia_Met | IPR041872 | Methionyl-tRNA synthetase, anticodon-binding domain | 373 | 505 | 5.0871E-33 |
| FunFam | G3DSA:2.20.28.20:FF:000001 | Methionine--tRNA ligase | - | - | 141 | 175 | 3.4E-16 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 545 | 564 | - |
| Gene3D | G3DSA:2.20.28.20 | - | IPR029038 | Methionyl-tRNA synthetase, Zn-domain | 141 | 175 | 0.0 |
| Gene3D | G3DSA:1.10.730.10 | - | - | - | 388 | 526 | 5.2E-35 |
| FunFam | G3DSA:2.40.50.140:FF:000042 | Methionine--tRNA ligase | - | - | 571 | 681 | 7.1E-44 |
| Pfam | PF19303 | Anticodon binding domain of methionyl tRNA ligase | IPR041872 | Methionyl-tRNA synthetase, anticodon-binding domain | 416 | 517 | 2.8E-6 |
| Pfam | PF01588 | Putative tRNA binding domain | IPR002547 | tRNA-binding domain | 585 | 678 | 5.5E-23 |
| PRINTS | PR01041 | Methionyl-tRNA synthetase signature | IPR033911 | Methioninyl-tRNA synthetase core domain | 89 | 100 | 2.6E-24 |
| SUPERFAMILY | SSF57770 | Methionyl-tRNA synthetase (MetRS), Zn-domain | IPR029038 | Methionyl-tRNA synthetase, Zn-domain | 141 | 175 | 2.75E-12 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.