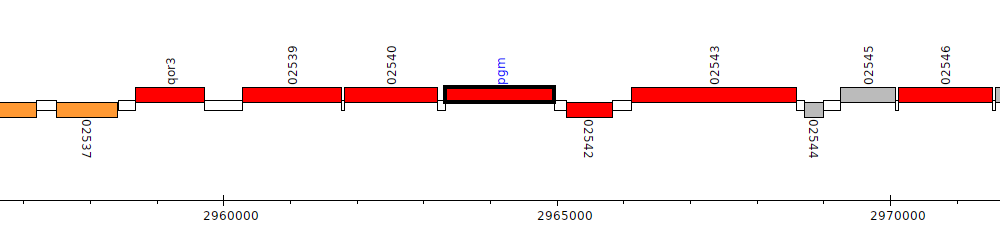
Pseudomonas sp. UW4, PputUW4_02541 (pgm)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004614 | phosphoglucomutase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01132
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0071704 | organic substance metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00408
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016868 | intramolecular transferase activity, phosphotransferases |
Inferred from Sequence Model
Term mapped from: InterPro:PF00408
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02879
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppuu01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | starch degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu00500 | Starch and sucrose metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | D-galactose degradation I (Leloir pathway) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | trehalose degradation V | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glucosylglycerol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | starch degradation V | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu00052 | Galactose metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | chitin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | sucrose biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glycogen biosynthesis III (from α-maltose 1-phosphate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | UDP-α-D-glucose biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | streptomycin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu00521 | Streptomycin biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | starch biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glycogen degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu00520 | Amino sugar and nucleotide sugar metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | CMP-legionaminate biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | sucrose degradation II (sucrose synthase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | ppuu00010 | Glycolysis / Gluconeogenesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppuu01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | GDP-glucose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | sucrose degradation IV (sucrose phosphorylase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 13 | 219 | 6.8E-63 |
| CDD | cd05801 | PGM_like3 | IPR005852 | Phosphoglucomutase, alpha-D-glucose specific | 20 | 541 | 0.0 |
| Pfam | PF02880 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain III | IPR005846 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain III | 323 | 439 | 2.2E-28 |
| NCBIfam | TIGR01132 | JCVI: phosphoglucomutase (alpha-D-glucose-1,6-bisphosphate-dependent) | IPR005852 | Phosphoglucomutase, alpha-D-glucose specific | 1 | 545 | 0.0 |
| Pfam | PF02878 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain I | IPR005844 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain I | 40 | 181 | 3.8E-34 |
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 222 | 319 | 5.6E-46 |
| Gene3D | G3DSA:3.40.120.10 | - | - | - | 320 | 443 | 2.8E-34 |
| SUPERFAMILY | SSF53738 | Phosphoglucomutase, first 3 domains | IPR016055 | Alpha-D-phosphohexomutase, alpha/beta/alpha I/II/III | 25 | 236 | 2.35E-48 |
| Pfam | PF02879 | Phosphoglucomutase/phosphomannomutase, alpha/beta/alpha domain II | IPR005845 | Alpha-D-phosphohexomutase, alpha/beta/alpha domain II | 211 | 318 | 4.2E-15 |
| SUPERFAMILY | SSF53738 | Phosphoglucomutase, first 3 domains | IPR016055 | Alpha-D-phosphohexomutase, alpha/beta/alpha I/II/III | 306 | 448 | 3.49E-32 |
| SUPERFAMILY | SSF55957 | Phosphoglucomutase, C-terminal domain | IPR036900 | Alpha-D-phosphohexomutase, C-terminal domain superfamily | 435 | 546 | 1.44E-20 |
| Gene3D | G3DSA:3.30.310.50 | - | - | - | 445 | 547 | 3.0E-39 |
| PANTHER | PTHR45745 | PHOSPHOMANNOMUTASE 45A | - | - | 19 | 542 | 1.2E-62 |
| Pfam | PF00408 | Phosphoglucomutase/phosphomannomutase, C-terminal domain | IPR005843 | Alpha-D-phosphohexomutase, C-terminal | 492 | 538 | 3.3E-10 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.