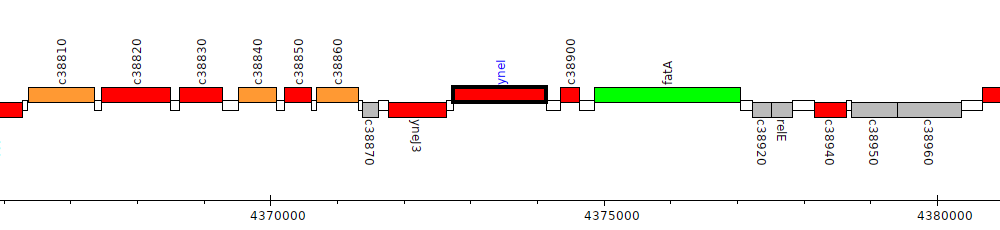
Pseudomonas protegens CHA0, PFLCHA0_c38890 (yneI)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53720
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.309.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004030 | aldehyde dehydrogenase [NAD(P)+] activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd07100
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pprc01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00650 | Butanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00250 | Alanine, aspartate and glutamate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00760 | Nicotinate and nicotinamide metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00171 | Aldehyde dehydrogenase family | IPR015590 | Aldehyde dehydrogenase domain | 10 | 458 | 0.0 |
| Gene3D | G3DSA:3.40.309.10 | Aldehyde Dehydrogenase; Chain A, domain 2 | IPR016163 | Aldehyde dehydrogenase, C-terminal | 238 | 427 | 0.0 |
| FunFam | G3DSA:3.40.309.10:FF:000010 | Gamma-aminobutyraldehyde dehydrogenase | - | - | 238 | 429 | 3.5E-71 |
| FunFam | G3DSA:3.40.605.10:FF:000012 | NAD-dependent succinate-semialdehyde dehydrogenase | - | - | 10 | 247 | 1.9E-80 |
| SUPERFAMILY | SSF53720 | ALDH-like | IPR016161 | Aldehyde/histidinol dehydrogenase | 7 | 461 | 0.0 |
| CDD | cd07100 | ALDH_SSADH1_GabD1 | IPR044148 | Succinate-semialdehyde dehydrogenase GabD1-like | 33 | 459 | 0.0 |
| Gene3D | G3DSA:3.40.605.10 | Aldehyde Dehydrogenase; Chain A, domain 1 | IPR016162 | Aldehyde dehydrogenase, N-terminal | 14 | 450 | 0.0 |
| PANTHER | PTHR43217 | SUCCINATE SEMIALDEHYDE DEHYDROGENASE [NAD(P)+] SAD | IPR047110 | Succinate-semialdehyde dehydrogenase [NADP(+)] GABD/Sad-like | 11 | 460 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.