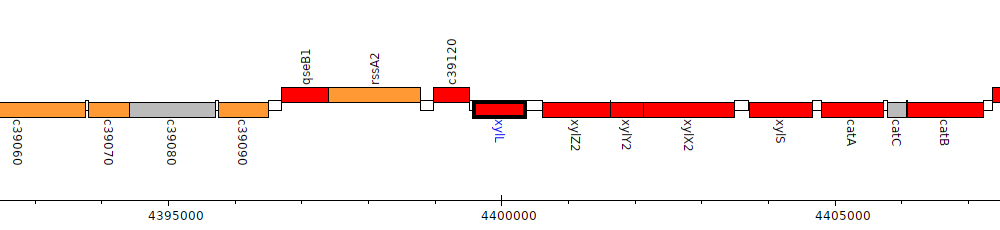
Pseudomonas protegens CHA0, PFLCHA0_c39130 (xylL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pprc01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00622 | Xylene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00362 | Benzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc00364 | Fluorobenzoate degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pprc01220 | Degradation of aromatic compounds | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 128 | 144 | 9.2E-38 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 179 | 202 | - |
| PANTHER | PTHR42760 | SHORT-CHAIN DEHYDROGENASES/REDUCTASES FAMILY MEMBER | - | - | 3 | 254 | 8.2E-55 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 1 | 256 | 9.4E-68 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 219 | 239 | 9.2E-38 |
| CDD | cd08937 | DHB_DH-like_SDR_c | - | - | 3 | 257 | 0.0 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 80 | 91 | 9.2E-38 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 8 | 25 | 9.2E-38 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 6 | 254 | 7.85E-67 |
| Pfam | PF00106 | short chain dehydrogenase | IPR002347 | Short-chain dehydrogenase/reductase SDR | 7 | 188 | 3.6E-50 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 152 | 171 | 9.2E-38 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 134 | 142 | 1.6E-10 |
| FunFam | G3DSA:3.40.50.720:FF:000084 | Short-chain dehydrogenase reductase | - | - | 1 | 256 | 5.1E-62 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 152 | 171 | 1.6E-10 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 80 | 91 | 1.6E-10 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 173 | 190 | 9.2E-38 |
| NCBIfam | NF040811 | NCBIFAM: benzoate diol dehydrogenase BenD | IPR047686 | Benzoate diol dehydrogenase BenD | 3 | 257 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.