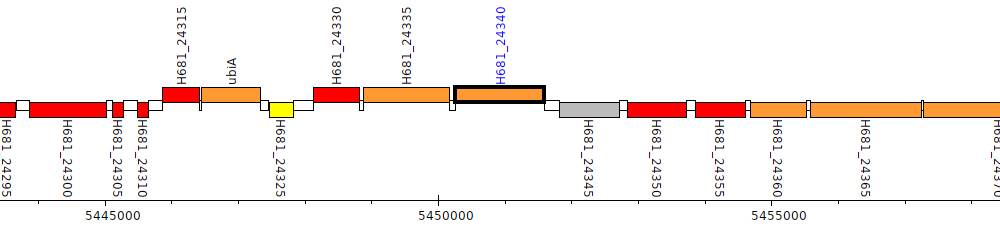
Pseudomonas denitrificans ATCC 13867, H681_24340
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF56176
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd04590 | CBS_pair_CorC_HlyC_assoc | IPR044751 | Ion transporter-like, CBS domain | 237 | 350 | 6.24199E-32 |
| SMART | SM00116 | cbs_1 | IPR000644 | CBS domain | 239 | 288 | 9.8E-4 |
| SMART | SM00116 | cbs_1 | IPR000644 | CBS domain | 305 | 353 | 0.049 |
| PANTHER | PTHR43099 | UPF0053 PROTEIN YRKA | - | - | 15 | 444 | 0.0 |
| Gene3D | G3DSA:3.10.580.10 | - | IPR046342 | CBS domain superfamily | 213 | 358 | 4.1E-34 |
| Pfam | PF03471 | Transporter associated domain | IPR005170 | Transporter-associated domain | 371 | 444 | 1.0E-15 |
| Pfam | PF00571 | CBS domain | IPR000644 | CBS domain | 302 | 353 | 1.1E-5 |
| SUPERFAMILY | SSF56176 | FAD-binding/transporter-associated domain-like | IPR036318 | FAD-binding, type PCMH-like superfamily | 367 | 445 | 1.25E-14 |
| Pfam | PF00571 | CBS domain | IPR000644 | CBS domain | 237 | 287 | 3.2E-8 |
| SMART | SM01091 | CorC_HlyC_2 | IPR005170 | Transporter-associated domain | 368 | 446 | 1.3E-16 |
| Gene3D | G3DSA:3.30.465.10 | - | IPR016169 | FAD-binding, type PCMH, subdomain 2 | 359 | 445 | 1.5E-15 |
| SUPERFAMILY | SSF54631 | CBS-domain pair | IPR046342 | CBS domain superfamily | 227 | 369 | 1.64E-25 |
| Pfam | PF01595 | Cyclin M transmembrane N-terminal domain | IPR002550 | CNNM, transmembrane domain | 21 | 209 | 8.1E-53 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.