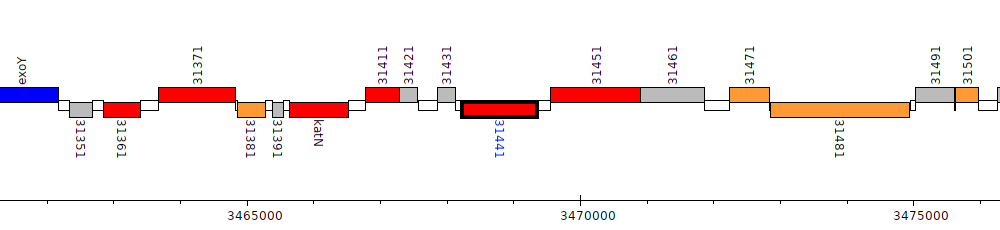
Pseudomonas aeruginosa LESB58, PALES_31441
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55931
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042398 | cellular modified amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF04107
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004357 | glutamate-cysteine ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF04107
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016879 | ligase activity, forming carbon-nitrogen bonds |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02050
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF55931 | Glutamine synthetase/guanido kinase | IPR014746 | Glutamine synthetase/guanido kinase, catalytic domain | 8 | 369 | 5.69E-87 |
| FunFam | G3DSA:3.30.590.20:FF:000008 | Putative glutamate--cysteine ligase 2 | - | - | 6 | 370 | 0.0 |
| NCBIfam | TIGR02050 | JCVI: YbdK family carboxylate-amine ligase | IPR011793 | Putative glutamate--cysteine ligase YbdK | 11 | 295 | 7.9E-115 |
| PANTHER | PTHR36510 | GLUTAMATE--CYSTEINE LIGASE 2-RELATED | - | - | 6 | 372 | 1.9E-107 |
| Hamap | MF_01609 | Glutamate--cysteine ligase [gshA]. | IPR011793 | Putative glutamate--cysteine ligase YbdK | 1 | 377 | 74.448654 |
| Pfam | PF04107 | Glutamate-cysteine ligase family 2(GCS2) | IPR006336 | Glutamate--cysteine ligase, GCS2 | 12 | 290 | 4.5E-89 |
| Gene3D | G3DSA:3.30.590.20 | - | - | - | 6 | 370 | 7.9E-87 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.