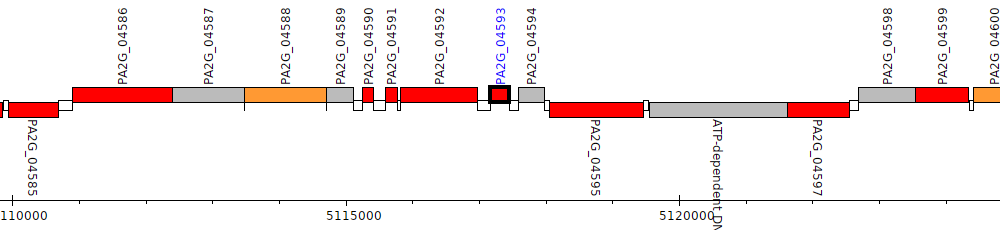
Pseudomonas aeruginosa 2192, PA2G_04593
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0030527 | structural constituent of chromatin |
Inferred from Sequence Model
Term mapped from: InterPro:PF00216
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47729
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00216 | Bacterial DNA-binding protein | IPR000119 | Histone-like DNA-binding protein | 1 | 90 | 1.5E-32 |
| Gene3D | G3DSA:4.10.520.10 | - | IPR010992 | Integration host factor (IHF)-like DNA-binding domain superfamily | 1 | 90 | 3.9E-32 |
| PRINTS | PR01727 | Prokaryotic integration host factor signature | IPR000119 | Histone-like DNA-binding protein | 40 | 55 | 2.1E-16 |
| SUPERFAMILY | SSF47729 | IHF-like DNA-binding proteins | IPR010992 | Integration host factor (IHF)-like DNA-binding domain superfamily | 1 | 90 | 2.65E-31 |
| PANTHER | PTHR33175 | DNA-BINDING PROTEIN HU | IPR000119 | Histone-like DNA-binding protein | 2 | 89 | 2.1E-26 |
| Coils | Coil | Coil | - | - | 11 | 38 | - |
| SMART | SM00411 | bhlneu | IPR000119 | Histone-like DNA-binding protein | 1 | 90 | 3.5E-38 |
| CDD | cd13831 | HU | - | - | 3 | 87 | 5.6708E-40 |
| PRINTS | PR01727 | Prokaryotic integration host factor signature | IPR000119 | Histone-like DNA-binding protein | 58 | 71 | 2.1E-16 |
| PRINTS | PR01727 | Prokaryotic integration host factor signature | IPR000119 | Histone-like DNA-binding protein | 74 | 88 | 2.1E-16 |
| FunFam | G3DSA:4.10.520.10:FF:000004 | HU family DNA-binding protein | - | - | 1 | 90 | 2.7E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.