Pseudomonas aeruginosa M18, PAM18_1222 (lasB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
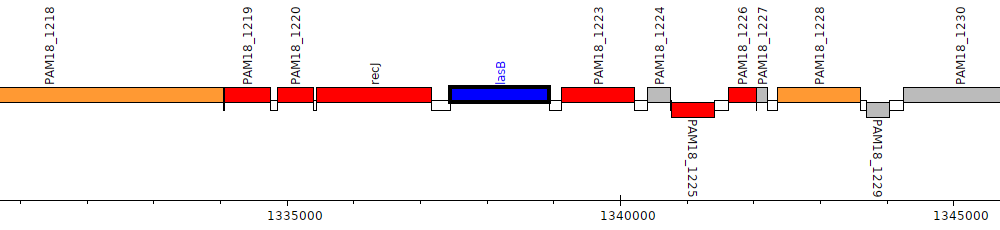
Gene Feature Overview
| Strain |
Pseudomonas aeruginosa M18
GCF_000226155.1|latest |
| Locus Tag |
PAM18_1222
|
| Name |
lasB
|
| Replicon | chromosome |
| Genomic location | 1337443 - 1338939 (+ strand) |
Cross-References
| RefSeq | YP_005973811.1 |
| GI | 386057289 |
| Entrez | 12565251 |
| NCBI Locus Tag | PAM18_1222 |
Product
| Feature Type | CDS |
| Coding Frame | 1 |
| Product Name |
elastase LasB
|
| Synonyms | |
| Evidence for Translation | |
| Charge (pH 7) | -1.48 |
| Kyte-Doolittle Hydrophobicity Value | -0.271 |
| Molecular Weight (kDa) | 53.7 |
| Isoelectric Point (pI) | 6.74 |
Subcellular localization
| Individual Mappings | |
| Additional evidence for subcellular localization |
AlphaFold 2 Protein Structure Predictions
Protein structure predictions using a neural network model developed by DeepMind. If a UniProtKB accession is associated with this protein, a search link will be provided below.
PDB 3D Structures
| Accession | Header | Accession Date | Compound | Source | Resolution | Method | Percent Identity |
| 7AJR | HYDROLASE | 09/29/20 | Virtual screening approach leading to the identification of a novel and tractable series of Pseudomonas aeruginosa elastase inhibitors | Pseudomonas aeruginosa | 1.75 | X-RAY DIFFRACTION | 100.0 |
| 3DBK | HYDROLASE | 06/01/08 | Pseudomonas aeruginosa elastase with phosphoramidon | Pseudomonas aeruginosa | 1.4 | X-RAY DIFFRACTION | 100.0 |
| 1EZM | HYDROLASE | 01/13/92 | THREE-DIMENSIONAL STRUCTURE OF THE ELASTASE OF PSEUDOMONAS AERUGINOSA AT 1.5 ANGSTROMS RESOLUTION | Pseudomonas aeruginosa | 1.5 | X-RAY DIFFRACTION | 100.0 |
| 8CR7 | HYDROLASE | 03/07/23 | Crystal structure of recombinant LasB from Pseudomonas aeruginosa PA7 | Pseudomonas aeruginosa PA7 | 1.5 | X-RAY DIFFRACTION | 95.9 |
| 7QH1 | HYDROLASE | 12/10/21 | Discovery and development of a novel inhaled antivirulence therapy for the treatment of Pseudomonas aeruginosa infections in patients with chronic respiratory disease | Pseudomonas aeruginosa | 2.74 | X-RAY DIFFRACTION | 100.0 |
| 1U4G | HYDROLASE | 07/25/04 | Elastase of Pseudomonas aeruginosa with an inhibitor | Pseudomonas aeruginosa | 1.4 | X-RAY DIFFRACTION | 100.0 |
| 6F8B | HYDROLASE | 12/12/17 | LasB bound to thiol based inhibitor | Pseudomonas aeruginosa | 1.3 | X-RAY DIFFRACTION | 100.0 |
| 7OC7 | HYDROLASE | 04/26/21 | LasB, alpha-alkyl-N-aryl mercaptoacetamide | Pseudomonas aeruginosa | 1.95 | X-RAY DIFFRACTION | 98.7 |
| 6FZX | HYDROLASE | 03/15/18 | LasB, hydroxymate Inhibitor Complex | Pseudomonas aeruginosa | 2.1 | X-RAY DIFFRACTION | 100.0 |
| 4K89 | HYDROLASE | 04/18/13 | Crystal structure of Pseudomonas aeruginosa strain K solvent tolerant elastase | Pseudomonas aeruginosa | 1.39 | X-RAY DIFFRACTION | 99.7 |
| 8CR3 | HYDROLASE | 03/07/23 | Crystal structure of recombinant LasB from Pseudomonas aeruginosa PAO1 | Pseudomonas aeruginosa PAO1 | 1.12 | X-RAY DIFFRACTION | 98.4 |
| 7NLK | HYDROLASE | 02/22/21 | LasB, N-aryl-2-butylmercaptoacetamide | Pseudomonas aeruginosa (strain UCBPP-PA14) | 1.7 | X-RAY DIFFRACTION | 98.7 |
| 7NLM | HYDROLASE | 02/22/21 | LasB, N-aryl-2-butylmercaptoacetamide | Pseudomonas aeruginosa (strain UCBPP-PA14) | 1.65 | X-RAY DIFFRACTION | 98.7 |
| 8CR4 | HYDROLASE | 03/07/23 | Crystal structure of recombinant LasBArtif from Pseudomonas aeruginosa AZPAE14816 | Pseudomonas aeruginosa | 0.91 | X-RAY DIFFRACTION | 94.9 |
Drugs Targeting this Protein
Identified by Diamond using e-value cutoff of 0.0001 and returning alignments that span 100% of the query sequence and that have more than 95% identity.
| Drug Name | Source Accession | Source DB | Version | Target Accession | Target Description | Percent Identity | Alignment Length | E-Value |
| N-(1-Carboxy-3-Phenylpropyl)Phenylalanyl-Alpha-Asparagine | DB02307 | DrugBank | 5.1.4 | P14756 | Elastase | 100.0 | 498 | 6.5e-289 |
Pathogen Association Analysis
| Results |
Common
Found in both pathogen and nonpathogenic strains
Hits to this gene were found in 114 genera
|
Orthologs/Comparative Genomics
| Pseudomonas Ortholog Database | View orthologs at Pseudomonas Ortholog Database |
| Pseudomonas Ortholog Group |
POG001959 (310 members) |
| Putative Inparalogs | None Found |
Interactions
| STRING database | Search for predicted protein-protein interactions using:
Search term: PAM18_1222
Search term: lasB
Search term: elastase LasB
|
Human Homologs
References
No references are associated with this feature.