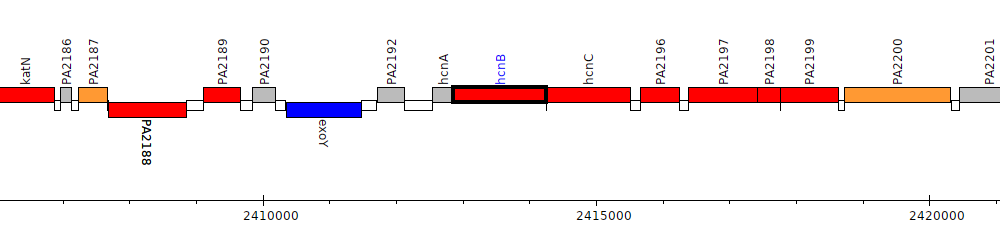
Pseudomonas aeruginosa PAO1, PA2194 (hcnB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008152 | metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF037495
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00460 | Cyanoamino acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 6 | 329 | 9.5E-22 |
| Pfam | PF04324 | BFD-like [2Fe-2S] binding domain | IPR007419 | BFD-like [2Fe-2S]-binding domain | 381 | 424 | 9.8E-12 |
| PANTHER | PTHR42949 | ANAEROBIC GLYCEROL-3-PHOSPHATE DEHYDROGENASE SUBUNIT B | - | - | 2 | 438 | 4.5E-116 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 120 | 264 | 9.5E-22 |
| CDD | cd19946 | GlpA-like_Fer2_BFD-like | - | - | 380 | 433 | 1.61912E-20 |
| PIRSF | PIRSF037495 | OoxA/HcnB | IPR017224 | Opine oxidase, subunit A/hydrogen cyanide synthase, subunit B | 1 | 464 | 0.0 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 6 | 311 | 3.7E-21 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 257 | 273 | 1.8E-10 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 107 | 125 | 1.8E-10 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 288 | 310 | 1.8E-10 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 6 | 330 | 3.96E-30 |
| Gene3D | G3DSA:1.10.10.1100 | - | IPR041854 | BFD-like [2Fe-2S]-binding domain superfamily | 369 | 439 | 1.3E-18 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 5 | 24 | 1.8E-10 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.