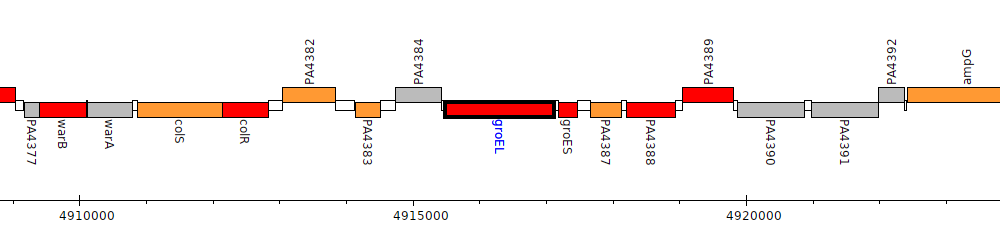
Pseudomonas aeruginosa PAO1, PA4385 (groEL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Operons
|
Operon name:
groESL
(2
gene members)
Experiment
Attributes
method:
S1 nuclease mapping, Northern blot
Evidence
Transcription unit experimentally validated in a strain of the same species. The start coordinate will often be based on an experimentally validated transcription start site. For protein-coding genes, unless otherwise specified, the stop will be based on the stop codon.
References
Transcription of the groESL operon in Pseudomonas aeruginosa PAO1.
Fujita M, Amemura A, Aramaki H
FEMS Microbiol. Lett. 1998 Jun;163(2):237-42
PubMed ID: 9673028
Cross-References
PseudoCAP:
41707
|
||||||||||||||||||||||||
|
Operon name:
groES-groEL
(2
gene members)
Evidence
Computationally-predicted.
References
DOOR: a database for prokaryotic operons.
Mao F, Dam P, Chou J, Olman V, Xu Y
Nucleic Acids Res. 2009 Jan;37(Database issue):D459-63
PubMed ID: 18988623
Cross-References
DOOR:
14002
|
||||||||||||||||||||||||
|
Operon name:
groESL
(2
gene members)
Experiment
Attributes
method:
S1 nuclease mapping, Northern blot
Evidence
Transcription unit experimentally validated in a strain of the same species. The start coordinate will often be based on an experimentally validated transcription start site. For protein-coding genes, unless otherwise specified, the stop will be based on the stop codon.
References
Transcription of the groESL operon in Pseudomonas aeruginosa PAO1.
Fujita M, Amemura A, Aramaki H
FEMS Microbiol. Lett. 1998 Jun;163(2):237-42
PubMed ID: 9673028
Cross-References
PseudoCAP:
41706
|