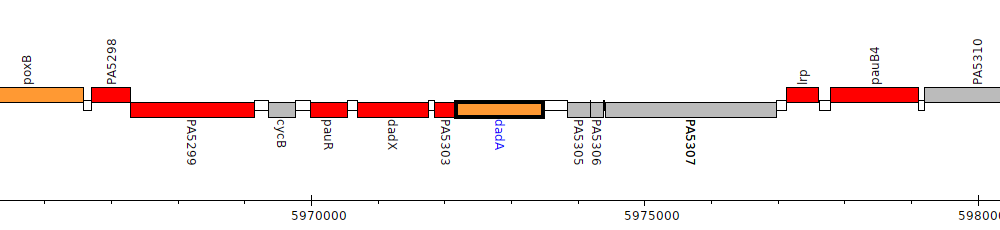
Pseudomonas aeruginosa PAO1, PA5304 (dadA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006807 | nitrogen compound metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0008718 | D-amino-acid dehydrogenase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
21378189 | Reviewed by curator |
| Biological Process | GO:0006558 | L-phenylalanine metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004795 | threonine synthase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0019478 | D-amino acid catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01202
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008718 | D-amino-acid dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01202
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01266
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00360 | Phenylalanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-6422 | D-arginine degradation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Phenylalanine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Nitrogen metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF54373 | FAD-linked reductases, C-terminal domain | - | - | 270 | 360 | 1.11E-17 |
| Hamap | MF_01202 | D-amino acid dehydrogenase [dadA]. | IPR023080 | D-amino acid dehydrogenase DadA | 1 | 411 | 50.179291 |
| Gene3D | G3DSA:3.30.9.10 | - | - | - | 134 | 363 | 3.7E-52 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 410 | 424 | - |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 416 | 2.07E-54 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 102 | 405 | 3.7E-52 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 1 | 71 | 2.5E-9 |
| PANTHER | PTHR13847 | SARCOSINE DEHYDROGENASE-RELATED | - | - | 2 | 419 | 4.6E-53 |
| FunFam | G3DSA:3.50.50.60:FF:000020 | D-amino acid dehydrogenase | - | - | 3 | 62 | 2.9E-34 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 410 | 432 | - |
| Pfam | PF01266 | FAD dependent oxidoreductase | IPR006076 | FAD dependent oxidoreductase | 3 | 398 | 1.7E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.